Identification of fusion genes in breast cancer by paired-end RNA-sequencing
- PMID: 21247443
- PMCID: PMC3091304
- DOI: 10.1186/gb-2011-12-1-r6
Identification of fusion genes in breast cancer by paired-end RNA-sequencing
Abstract
Background: Until recently, chromosomal translocations and fusion genes have been an underappreciated class of mutations in solid tumors. Next-generation sequencing technologies provide an opportunity for systematic characterization of cancer cell transcriptomes, including the discovery of expressed fusion genes resulting from underlying genomic rearrangements.
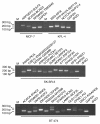
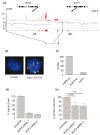
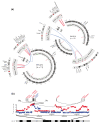
Results: We applied paired-end RNA-seq to identify 24 novel and 3 previously known fusion genes in breast cancer cells. Supported by an improved bioinformatic approach, we had a 95% success rate of validating gene fusions initially detected by RNA-seq. Fusion partner genes were found to contribute promoters (5' UTR), coding sequences and 3' UTRs. Most fusion genes were associated with copy number transitions and were particularly common in high-level DNA amplifications. This suggests that fusion events may contribute to the selective advantage provided by DNA amplifications and deletions. Some of the fusion partner genes, such as GSDMB in the TATDN1-GSDMB fusion and IKZF3 in the VAPB-IKZF3 fusion, were only detected as a fusion transcript, indicating activation of a dormant gene by the fusion event. A number of fusion gene partners have either been previously observed in oncogenic gene fusions, mostly in leukemias, or otherwise reported to be oncogenic. RNA interference-mediated knock-down of the VAPB-IKZF3 fusion gene indicated that it may be necessary for cancer cell growth and survival.
Conclusions: In summary, using RNA-sequencing and improved bioinformatic stratification, we have discovered a number of novel fusion genes in breast cancer, and identified VAPB-IKZF3 as a potential fusion gene with importance for the growth and survival of breast cancer cells.
Figures


References
-
- Tomlins SA, Rhodes DR, Perner S, Dhanasekaran SM, Mehra R, Sun XW, Varambally S, Cao X, Tchinda J, Kuefer R, Lee C, Montie JE, Shah RB, Pienta KJ, Rubin MA, Chinnaiyan AM. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science. 2005;310:644–648. doi: 10.1126/science.1117679. - DOI - PubMed
-
- Soda M, Choi YL, Enomoto M, Takada S, Yamashita Y, Ishikawa S, Fujiwara S, Watanabe H, Kurashina K, Hatanaka H, Bando M, Ohno S, Ishikawa Y, Aburatani H, Niki T, Sohara Y, Sugiyama Y, Mano H. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature. 2007;448:561–566. doi: 10.1038/nature05945. - DOI - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

