Analysis of human APOBEC3H haplotypes and anti-human immunodeficiency virus type 1 activity
- PMID: 21270145
- PMCID: PMC3067873
- DOI: 10.1128/JVI.02049-10
Analysis of human APOBEC3H haplotypes and anti-human immunodeficiency virus type 1 activity
Abstract
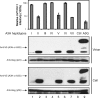
Human APOBEC3H (A3H) has one cytidine deaminase domain (CDD) and inhibits the replication of retrotransposons and human immunodeficiency virus type 1 (HIV-1) in a Vif-resistant manner. Human A3H has five single amino acid polymorphisms (N15Δ, R18L, G105R, K121D, and E178D), and four haplotypes (I to IV) have previously been identified in various human populations. Haplotype II was primarily found in African-derived populations, and it was the only one that could be stably expressed. Here, we identified three new haplotypes from six human population samples, which we have named V, VI, and VII. Haplotypes V and VII are stably expressed and inhibit HIV-1 replication. Notably, haplotype V was identified in samples from all African-, Asian-, and Caucasian-derived populations studied. Using haplotype VII, we investigated the A3H anti-HIV-1 mechanism. We found that A3H virion packaging is independent of its CDD but dependent on a (112)YYXW(115) motif. This motif binds HIV-1 nucleocapsid in an RNA-dependent manner, and a single Y112A mutation completely disrupts A3H virion incorporation. We further studied the mechanism of A3H resistance to Vif. Although the previously identified APOBEC3G Vif-responsive motif (128)DPDY(131) is not conserved in A3H, placement of this motif into A3H does not make it become less resistant to HIV-1 Vif. We conclude that stably expressed A3H haplotypes may be more broadly distributed in humans than previously realized, and A3H protein is resistant to Vif. These results have important implications for the role of A3H in retrotransposon and HIV-1 inhibition.
Figures

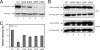



References
-
- Bishop, K. N., et al. 2004. Cytidine deamination of retroviral DNA by diverse APOBEC proteins. Curr. Biol. 14:1392-1396. - PubMed
-
- Clark, A. G. 1990. Inference of haplotypes from PCR-amplified samples of diploid populations. Mol. Biol. Evol. 7:111-122. - PubMed
-
- Daly, M. J., J. D. Rioux, S. F. Schaffner, T. J. Hudson, and E. S. Lander. 2001. High-resolution haplotype structure in the human genome. Nat. Genet. 29:229-232. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

