CaMtw1, a member of the evolutionarily conserved Mis12 kinetochore protein family, is required for efficient inner kinetochore assembly in the pathogenic yeast Candida albicans
- PMID: 21276093
- PMCID: PMC3086999
- DOI: 10.1111/j.1365-2958.2011.07558.x
CaMtw1, a member of the evolutionarily conserved Mis12 kinetochore protein family, is required for efficient inner kinetochore assembly in the pathogenic yeast Candida albicans
Abstract
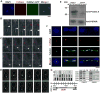
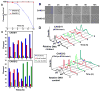
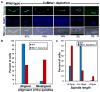
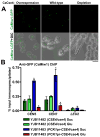
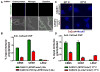
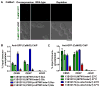
Proper assembly of the kinetochore, a multi-protein complex that mediates attachment of centromere DNA to spindle microtubules on each chromosome, is required for faithful chromosome segregation. Each previously characterized member of the Mis12/Mtw1 protein family is part of an essential subcomplex in the kinetochore. In this work, we identify and characterize CaMTW1, which encodes the homologue of the human Mis12 protein in the pathogenic budding yeast Candida albicans. Subcellular localization and chromatin immunoprecipitation assays confirmed CaMtw1 is a kinetochore protein. CaMtw1 is essential for viability. CaMtw1-depleted cells and cells in which CaMtw1 was inactivated with a temperature-sensitive mutation had reduced viability, accumulated at the G2/M stage of the cell cycle, and exhibited increased chromosome missegregation. CaMtw1 depletion also affected spindle length and alignment. Interestingly, in C. albicans, CaMtw1 and the centromeric histone, CaCse4, influence each other for kinetochore localization. In addition, CaMtw1 is required for efficient kinetochore recruitment of another inner kinetochore protein, the CENP-C homologue, CaMif2. Mis12/Mtw1 proteins have well-established roles in the recruitment and maintenance of outer kinetochore proteins. We propose that Mis12/Mtw1 proteins also have important co-dependent interactions with inner kinetochore proteins and that these interactions may increase the fidelity of kinetochore formation.
© 2011 Blackwell Publishing Ltd.
Figures


Similar articles
-
The essentiality of the fungus-specific Dam1 complex is correlated with a one-kinetochore-one-microtubule interaction present throughout the cell cycle, independent of the nature of a centromere.Eukaryot Cell. 2011 Oct;10(10):1295-305. doi: 10.1128/EC.05093-11. Epub 2011 May 13. Eukaryot Cell. 2011. PMID: 21571923 Free PMC article.
-
The CENP-A homolog CaCse4p in the pathogenic yeast Candida albicans is a centromere protein essential for chromosome transmission.Proc Natl Acad Sci U S A. 2002 Oct 1;99(20):12969-74. doi: 10.1073/pnas.162488299. Epub 2002 Sep 23. Proc Natl Acad Sci U S A. 2002. PMID: 12271118 Free PMC article.
-
A coordinated interdependent protein circuitry stabilizes the kinetochore ensemble to protect CENP-A in the human pathogenic yeast Candida albicans.PLoS Genet. 2012;8(4):e1002661. doi: 10.1371/journal.pgen.1002661. Epub 2012 Apr 19. PLoS Genet. 2012. PMID: 22536162 Free PMC article.
-
The kinetochore interaction network (KIN) of ascomycetes.Mycologia. 2016 May-Jun;108(3):485-505. doi: 10.3852/15-182. Epub 2016 Feb 23. Mycologia. 2016. PMID: 26908646 Free PMC article. Review.
-
Cell cycle control of kinetochore assembly.Nucleus. 2022 Dec;13(1):208-220. doi: 10.1080/19491034.2022.2115246. Nucleus. 2022. PMID: 36037227 Free PMC article. Review.
Cited by
-
A surprising role for the Sch9 protein kinase in chromosome segregation in Candida albicans.Genetics. 2015 Mar;199(3):671-4. doi: 10.1534/genetics.114.173542. Epub 2015 Jan 15. Genetics. 2015. PMID: 25591453 Free PMC article.
-
The essentiality of the fungus-specific Dam1 complex is correlated with a one-kinetochore-one-microtubule interaction present throughout the cell cycle, independent of the nature of a centromere.Eukaryot Cell. 2011 Oct;10(10):1295-305. doi: 10.1128/EC.05093-11. Epub 2011 May 13. Eukaryot Cell. 2011. PMID: 21571923 Free PMC article.
-
Flexibility of centromere and kinetochore structures.Trends Genet. 2012 May;28(5):204-12. doi: 10.1016/j.tig.2012.02.003. Epub 2012 Mar 23. Trends Genet. 2012. PMID: 22445183 Free PMC article. Review.
-
Functional genomic analysis of genes important for Candida albicans fitness in diverse environmental conditions.Cell Rep. 2024 Aug 27;43(8):114601. doi: 10.1016/j.celrep.2024.114601. Epub 2024 Aug 8. Cell Rep. 2024. PMID: 39126650 Free PMC article.
-
Leveraging machine learning essentiality predictions and chemogenomic interactions to identify antifungal targets.Nat Commun. 2021 Nov 11;12(1):6497. doi: 10.1038/s41467-021-26850-3. Nat Commun. 2021. PMID: 34764269 Free PMC article.
References
-
- Bachewich C, Nantel A, Whiteway M. Cell cycle arrest during S or M phase generates polarized growth via distinct signals in Candida albicans. Mol Microbiol. 2005;57:942–959. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

