Structural basis of the 9-fold symmetry of centrioles
- PMID: 21277013
- PMCID: PMC3089914
- DOI: 10.1016/j.cell.2011.01.008
Structural basis of the 9-fold symmetry of centrioles
Abstract
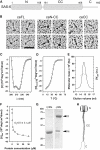
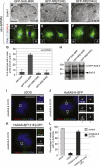
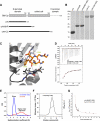
The centriole, and the related basal body, is an ancient organelle characterized by a universal 9-fold radial symmetry and is critical for generating cilia, flagella, and centrosomes. The mechanisms directing centriole formation are incompletely understood and represent a fundamental open question in biology. Here, we demonstrate that the centriolar protein SAS-6 forms rod-shaped homodimers that interact through their N-terminal domains to form oligomers. We establish that such oligomerization is essential for centriole formation in C. elegans and human cells. We further generate a structural model of the related protein Bld12p from C. reinhardtii, in which nine homodimers assemble into a ring from which nine coiled-coil rods radiate outward. Moreover, we demonstrate that recombinant Bld12p self-assembles into structures akin to the central hub of the cartwheel, which serves as a scaffold for centriole formation. Overall, our findings establish a structural basis for the universal 9-fold symmetry of centrioles.
Copyright © 2011 Elsevier Inc. All rights reserved.
Figures


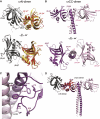








Comment in
-
Cytoskeleton: SAS-6 turns a cartwheel trick.Nat Rev Mol Cell Biol. 2011 Mar;12(3):137. doi: 10.1038/nrm3071. Nat Rev Mol Cell Biol. 2011. PMID: 21346728 No abstract available.
References
-
- Bach M., Grigat S., Pawlik B., Fork C., Utermöhlen O., Pal S., Banczyk D., Lazar A., Schömig E., Gründemann D. Fast set-up of doxycycline-inducible protein expression in human cell lines with a single plasmid based on Epstein-Barr virus replication and the simple tetracycline repressor. FEBS J. 2007;274:783–790. - PubMed
-
- Carvalho-Santos Z., Machado P., Branco P., Tavares-Cadete F., Rodrigues-Martins A., Pereira-Leal J.B., Bettencourt-Dias M. Stepwise evolution of the centriole-assembly pathway. J. Cell Sci. 2010;123:1414–1426. - PubMed
-
- Cavalier-Smith T. Basal body and flagellar development during the vegetative cell cycle and the sexual cycle of Chlamydomonas reinhardii. J. Cell Sci. 1974;16:529–556. - PubMed
Supplemental References
-
- Adams, P.D., Grosse-Kunstleve, R.W., Hung, L.W., Ioerger, T.R., McCoy, A.J., Moriarty, N.W., Read, R.J., Sacchettini, J.C., Sauter, N.K., and Terwilliger, T.C. (2002). PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954. - PubMed
-
- Bach, M., Grigat, S., Pawlik, B., Fork, C., Utermöhlen, O., Pal, S., Banczyk, D., Lazar, A., Schömig, E., and Gründemann, D. (2007). Fast set-up of doxycycline-inducible protein expression in human cell lines with a single plasmid based on Epstein-Barr virus replication and the simple tetracycline repressor. FEBS J. 274, 783–790. - PubMed
-
- Bingel-Erlenmeyer, R., Olieric, V., Grimshaw, J.P.A., Gabadinho, J., Wang, X., Ebner, S.G., Isenegger, A., Schneider, R., Schneider, J., Glettig, W., et al. The SLS crystallization platform at beam-line X06DA - a fully automated pipeline enabling in situ X-ray diffraction screening. Cryst. Growth Des., in press.
-
- Blanc, E., Roversi, P., Vonrhein, C., Flensburg, C., Lea, S.M., and Bricogne, G. (2004). Refinement of severely incomplete structures with maximum likelihood in BUSTER-TNT. Acta Crystallogr. D Biol. Crystallogr. 60, 2210–2221. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

