PDE4D and PDE4B function in distinct subcellular compartments in mouse embryonic fibroblasts
- PMID: 21288894
- PMCID: PMC3069460
- DOI: 10.1074/jbc.M110.203604
PDE4D and PDE4B function in distinct subcellular compartments in mouse embryonic fibroblasts
Abstract
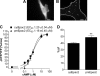
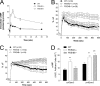
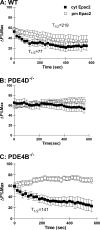
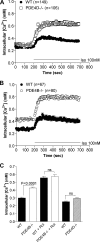
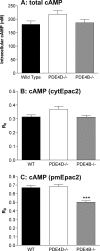
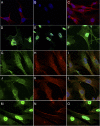
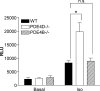
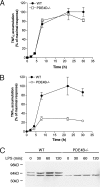
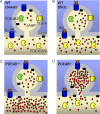
Signaling through cAMP regulates most cellular functions. The spatiotemporal control of cAMP is, therefore, crucial for differential regulation of specific cellular targets. Here we investigated the consequences of PDE4B or PDE4D gene ablation on cAMP signaling at a subcellular level using mouse embryonic fibroblasts. PDE4B ablation had no effect on the global or bulk cytosol accumulation of cAMP but increased both basal and hormone-dependent cAMP in a near-membrane pool. Conversely, PDE4D ablation enhanced agonist-induced cAMP accumulation in the bulk cytosol as well as at the plasma membrane. Both PDE4B and PDE4D ablation significantly modified the time course and the level of isoproterenol-induced phosphorylation of vasodilator-stimulated phosphoprotein, a membrane cytoskeletal component. A second membrane response through Toll-like receptor signaling, however, was only affected by PDE4B ablation. PDE4D but not PDE4B ablation significantly prolonged cAMP-response element-binding protein-mediated transcription. These findings demonstrate that PDE4D and PDE4B have specialized functions in mouse embryonic fibroblasts with PDE4B controlling cAMP in a discrete subdomain near the plasma membrane.
Figures


Similar articles
-
Critical role of PDE4D in beta2-adrenoceptor-dependent cAMP signaling in mouse embryonic fibroblasts.J Biol Chem. 2008 Aug 15;283(33):22430-42. doi: 10.1074/jbc.M803306200. Epub 2008 May 28. J Biol Chem. 2008. PMID: 18508768 Free PMC article.
-
Spatiotemporal dynamics of beta-adrenergic cAMP signals and L-type Ca2+ channel regulation in adult rat ventricular myocytes: role of phosphodiesterases.Circ Res. 2008 May 9;102(9):1091-100. doi: 10.1161/CIRCRESAHA.107.167817. Epub 2008 Mar 27. Circ Res. 2008. PMID: 18369156
-
Phosphodiesterases 4B and 4D Differentially Regulate cAMP Signaling in Calcium Handling Microdomains of Mouse Hearts.Cells. 2024 Mar 8;13(6):476. doi: 10.3390/cells13060476. Cells. 2024. PMID: 38534320 Free PMC article.
-
PDE4D phosphorylation: A coincidence detector integrating multiple signaling pathways.Cell Signal. 2016 Jul;28(7):719-24. doi: 10.1016/j.cellsig.2015.11.001. Epub 2015 Nov 10. Cell Signal. 2016. PMID: 26562185 Review.
-
Biophysical techniques for detection of cAMP and cGMP in living cells.Int J Mol Sci. 2013 Apr 12;14(4):8025-46. doi: 10.3390/ijms14048025. Int J Mol Sci. 2013. PMID: 23584022 Free PMC article. Review.
Cited by
-
Phosphodiesterase inhibitor modulation of brain microvascular endothelial cell barrier properties.J Neurol Sci. 2012 Sep 15;320(1-2):45-51. doi: 10.1016/j.jns.2012.06.005. Epub 2012 Jul 21. J Neurol Sci. 2012. PMID: 22819056 Free PMC article.
-
Ion Channel-Based Reporters for cAMP Detection.Methods Mol Biol. 2022;2483:265-279. doi: 10.1007/978-1-0716-2245-2_17. Methods Mol Biol. 2022. PMID: 35286682
-
Perspectives on: Cyclic nucleotide microdomains and signaling specificity.J Gen Physiol. 2014 Jan;143(1):5-7. doi: 10.1085/jgp.201311144. J Gen Physiol. 2014. PMID: 24378902 Free PMC article. No abstract available.
-
Assessment of cellular mechanisms contributing to cAMP compartmentalization in pulmonary microvascular endothelial cells.Am J Physiol Cell Physiol. 2012 Mar 15;302(6):C839-52. doi: 10.1152/ajpcell.00361.2011. Epub 2011 Nov 23. Am J Physiol Cell Physiol. 2012. PMID: 22116306 Free PMC article.
-
Effects of PDE4 pathway inhibition in rat experimental stroke.J Pharm Pharm Sci. 2014;17(3):362-70. doi: 10.18433/j3s02v. J Pharm Pharm Sci. 2014. PMID: 25224348 Free PMC article.
References
-
- Barnes A. P., Livera G., Huang P., Sun C., O'Neal W. K., Conti M., Stutts M. J., Milgram S. L. (2005) J. Biol. Chem. 280, 7997–8003 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

