Fused eco29kIR- and M genes coding for a fully functional hybrid polypeptide as a model of molecular evolution of restriction-modification systems
- PMID: 21291520
- PMCID: PMC3039593
- DOI: 10.1186/1471-2148-11-35
Fused eco29kIR- and M genes coding for a fully functional hybrid polypeptide as a model of molecular evolution of restriction-modification systems
Abstract
Background: The discovery of restriction endonucleases and modification DNA methyltransferases, key instruments of genetic engineering, opened a new era of molecular biology through development of the recombinant DNA technology. Today, the number of potential proteins assigned to type II restriction enzymes alone is beyond 6000, which probably reflects the high diversity of evolutionary pathways. Here we present experimental evidence that a new type IIC restriction and modification enzymes carrying both activities in a single polypeptide could result from fusion of the appropriate genes from preexisting bipartite restriction-modification systems.
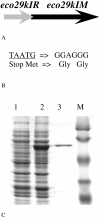
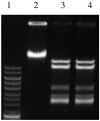
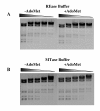
Results: Fusion of eco29kIR and M ORFs gave a novel gene encoding for a fully functional hybrid polypeptide that carried both restriction endonuclease and DNA methyltransferase activities. It has been placed into a subclass of type II restriction and modification enzymes--type IIC. Its MTase activity, 80% that of the M.Eco29kI enzyme, remained almost unchanged, while its REase activity decreased by three times, concurrently with changed reaction optima, which presumably can be caused by increased steric hindrance in interaction with the substrate. In vitro the enzyme preferentially cuts DNA, with only a low level of DNA modification detected. In vivo new RMS can provide a 102-fold less protection of host cells against phage invasion.
Conclusions: We propose a molecular mechanism of appearing of type IIC restriction-modification and M.SsoII-related enzymes, as well as other multifunctional proteins. As shown, gene fusion could play an important role in evolution of restriction-modification systems and be responsible for the enzyme subclass interconversion. Based on the proposed approach, hundreds of new type IIC enzymes can be generated using head-to-tail oriented type I, II, and III restriction and modification genes. These bifunctional polypeptides can serve a basis for enzymes with altered recognition specificities. Lastly, this study demonstrates that protein fusion may change biochemical properties of the involved enzymes, thus giving a starting point for their further evolutionary divergence.
Figures







Similar articles
-
Related bifunctional restriction endonuclease-methyltransferase triplets: TspDTI, Tth111II/TthHB27I and TsoI with distinct specificities.BMC Mol Biol. 2012 Apr 10;13:13. doi: 10.1186/1471-2199-13-13. BMC Mol Biol. 2012. PMID: 22489904 Free PMC article.
-
New bifunctional restriction-modification enzyme AloI isoschizomer (PcoI): Bioinformatics analysis, purification and activity confirmation.Gene. 2018 Jun 20;660:8-12. doi: 10.1016/j.gene.2018.03.069. Epub 2018 Mar 21. Gene. 2018. PMID: 29574188
-
Structure and evolution of the XcyI restriction-modification system.Nucleic Acids Res. 1992 Dec 11;20(23):6267-73. doi: 10.1093/nar/20.23.6267. Nucleic Acids Res. 1992. PMID: 1475187 Free PMC article.
-
Cloning type-II restriction and modification genes.Gene. 1988 Dec 25;74(1):25-32. doi: 10.1016/0378-1119(88)90242-9. Gene. 1988. PMID: 3074013 Review.
-
Type II restriction endonucleases: structure and mechanism.Cell Mol Life Sci. 2005 Mar;62(6):685-707. doi: 10.1007/s00018-004-4513-1. Cell Mol Life Sci. 2005. PMID: 15770420 Free PMC article. Review.
Cited by
-
Naturally-occurring, dually-functional fusions between restriction endonucleases and regulatory proteins.BMC Evol Biol. 2013 Oct 2;13:218. doi: 10.1186/1471-2148-13-218. BMC Evol Biol. 2013. PMID: 24083337 Free PMC article.
-
The interplay of restriction-modification systems with mobile genetic elements and their prokaryotic hosts.Nucleic Acids Res. 2014;42(16):10618-31. doi: 10.1093/nar/gku734. Epub 2014 Aug 12. Nucleic Acids Res. 2014. PMID: 25120263 Free PMC article.
-
A model for the evolution of prokaryotic DNA restriction-modification systems based upon the structural malleability of Type I restriction-modification enzymes.Nucleic Acids Res. 2018 Sep 28;46(17):9067-9080. doi: 10.1093/nar/gky760. Nucleic Acids Res. 2018. PMID: 30165537 Free PMC article.
-
Removal of a frameshift between the hsdM and hsdS genes of the EcoKI Type IA DNA restriction and modification system produces a new type of system and links the different families of Type I systems.Nucleic Acids Res. 2012 Nov;40(21):10916-24. doi: 10.1093/nar/gks876. Epub 2012 Sep 23. Nucleic Acids Res. 2012. PMID: 23002145 Free PMC article.
-
Structure and operation of the DNA-translocating type I DNA restriction enzymes.Genes Dev. 2012 Jan 1;26(1):92-104. doi: 10.1101/gad.179085.111. Genes Dev. 2012. PMID: 22215814 Free PMC article.
References
-
- Skowronek KJ, Bujnicki JM. In: Industrial Enzymes: Structure, Function and Applications. Polaina J, MacCabe AP, editor. Chapter 21. Springer; 2007.
-
- Roberts RJ, Belfort M, Bestor T, Bhagwat AS, Bickle TA, Bitinaite J, Blumenthal RM, Degtyarev S, Dryden DT, Dybvig K. et al.A nomenclature for restriction enzymes, DNA methyltransferases, homing endonucleases and their genes. Nucleic Acids Res. 2003;31:1805–1812. doi: 10.1093/nar/gkg274. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

