RNA content in the nucleolus alters p53 acetylation via MYBBP1A
- PMID: 21297583
- PMCID: PMC3061030
- DOI: 10.1038/emboj.2011.23
RNA content in the nucleolus alters p53 acetylation via MYBBP1A
Abstract
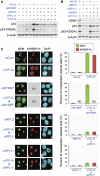
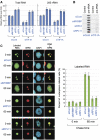
A number of external and internal insults disrupt nucleolar structure, and the resulting nucleolar stress stabilizes and activates p53. We show here that nucleolar disruption induces acetylation and accumulation of p53 without phosphorylation. We identified three nucleolar proteins, MYBBP1A, RPL5, and RPL11, involved in p53 acetylation and accumulation. MYBBP1A was tethered to the nucleolus through nucleolar RNA. When rRNA transcription was suppressed by nucleolar stress, MYBBP1A translocated to the nucleoplasm and facilitated p53-p300 interaction to enhance p53 acetylation. We also found that RPL5 and RPL11 were required for rRNA export from the nucleolus. Depletion of RPL5 or RPL11 blocked rRNA export and counteracted reduction of nucleolar RNA levels caused by inhibition of rRNA transcription. As a result, RPL5 or RPL11 depletion inhibited MYBBP1A translocation and p53 activation. Our observations indicated that a dynamic equilibrium between RNA generation and export regulated nucleolar RNA content. Perturbation of this balance by nucleolar stress altered the nucleolar RNA content and modulated p53 activity.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures




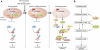
References
-
- Andersen JS, Lam YW, Leung AK, Ong SE, Lyon CE, Lamond AI, Mann M (2005) Nucleolar proteome dynamics. Nature 433: 77–83 - PubMed
-
- Appella E, Anderson CW (2001) Post-translational modifications and activation of p53 by genotoxic stresses. Eur J Biochem 268: 2764–2772 - PubMed
-
- Arva NC, Gopen TR, Talbott KE, Campbell LE, Chicas A, White DE, Bond GL, Levine AJ, Bargonetti J (2005) A chromatin-associated and transcriptionally inactive p53-Mdm2 complex occurs in mdm2 SNP309 homozygous cells. J Biol Chem 280: 26776–26787 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

