Positive signature-tagged mutagenesis in Pseudomonas aeruginosa: tracking patho-adaptive mutations promoting airways chronic infection
- PMID: 21304889
- PMCID: PMC3033382
- DOI: 10.1371/journal.ppat.1001270
Positive signature-tagged mutagenesis in Pseudomonas aeruginosa: tracking patho-adaptive mutations promoting airways chronic infection
Abstract
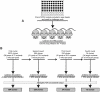
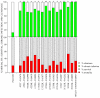
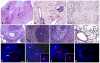
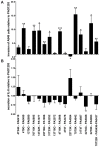
The opportunistic pathogen Pseudomonas aeruginosa can establish life-long chronic infections in the airways of cystic fibrosis (CF) patients. Persistent lifestyle is established with P. aeruginosa patho-adaptive variants, which are clonal with the initially-acquired strains. Several reports indicated that P. aeruginosa adapts by loss-of-function mutations which enhance fitness in CF airways and sustain its clonal expansion during chronic infection. To validate this model of P. aeruginosa adaptation to CF airways and to identify novel genes involved in this microevolution, we designed a novel approach of positive-selection screening by PCR-based signature-tagged mutagenesis (Pos-STM) in a murine model of chronic airways infection. A systematic positive-selection scheme using sequential rounds of in vivo screenings for bacterial maintenance, as opposed to elimination, generated a list of genes whose inactivation increased the colonization and persistence in chronic airways infection. The phenotypes associated to these Pos-STM mutations reflect alterations in diverse aspects of P. aeruginosa biology which include lack of swimming and twitching motility, lack of production of the virulence factors such as pyocyanin, biofilm formation, and metabolic functions. In addition, Pos-STM mutants showed altered invasion and stimulation of immune response when tested in human respiratory epithelial cells, indicating that P. aeruginosa is prone to revise the interaction with its host during persistent lifestyle. Finally, sequence analysis of Pos-STM genes in longitudinally P. aeruginosa isolates from CF patients identified signs of patho-adaptive mutations within the genome. This novel Pos-STM approach identified bacterial functions that can have important clinical implications for the persistent lifestyle and disease progression of the airway chronic infection.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
Similar articles
-
The Small RNA ErsA Plays a Role in the Regulatory Network of Pseudomonas aeruginosa Pathogenicity in Airway Infections.mSphere. 2020 Oct 14;5(5):e00909-20. doi: 10.1128/mSphere.00909-20. mSphere. 2020. PMID: 33055260 Free PMC article.
-
Pseudomonas aeruginosa microevolution during cystic fibrosis lung infection establishes clones with adapted virulence.Am J Respir Crit Care Med. 2009 Jul 15;180(2):138-45. doi: 10.1164/rccm.200812-1943OC. Epub 2009 May 7. Am J Respir Crit Care Med. 2009. PMID: 19423715
-
In vivo functional genomics of Pseudomonas aeruginosa for high-throughput screening of new virulence factors and antibacterial targets.Environ Microbiol. 2003 Dec;5(12):1294-308. doi: 10.1046/j.1462-2920.2003.00542.x. Environ Microbiol. 2003. PMID: 14641575
-
Microevolution of Pseudomonas aeruginosa to a chronic pathogen of the cystic fibrosis lung.Curr Top Microbiol Immunol. 2013;358:91-118. doi: 10.1007/82_2011_199. Curr Top Microbiol Immunol. 2013. PMID: 22311171 Review.
-
Intrinsic and environmental mutagenesis drive diversification and persistence of Pseudomonas aeruginosa in chronic lung infections.J Infect Dis. 2012 Jan 1;205(1):121-7. doi: 10.1093/infdis/jir690. Epub 2011 Nov 11. J Infect Dis. 2012. PMID: 22080096 Review.
Cited by
-
The YfiBNR signal transduction mechanism reveals novel targets for the evolution of persistent Pseudomonas aeruginosa in cystic fibrosis airways.PLoS Pathog. 2012;8(6):e1002760. doi: 10.1371/journal.ppat.1002760. Epub 2012 Jun 14. PLoS Pathog. 2012. PMID: 22719254 Free PMC article.
-
Efficient protocols and methods for high-throughput utilization of the Collaborative Cross mouse model for dissecting the genetic basis of complex traits.Animal Model Exp Med. 2019 Jul 10;2(3):137-149. doi: 10.1002/ame2.12074. eCollection 2019 Sep. Animal Model Exp Med. 2019. PMID: 31773089 Free PMC article. Review.
-
Selection for novel metabolic capabilities in Salmonella enterica.Evolution. 2019 May;73(5):990-1000. doi: 10.1111/evo.13713. Epub 2019 Mar 22. Evolution. 2019. PMID: 30848832 Free PMC article.
-
Adaptation of Pseudomonas aeruginosa in Cystic Fibrosis airways influences virulence of Staphylococcus aureus in vitro and murine models of co-infection.PLoS One. 2014 Mar 6;9(3):e89614. doi: 10.1371/journal.pone.0089614. eCollection 2014. PLoS One. 2014. PMID: 24603807 Free PMC article.
-
Mapping genetic determinants of host susceptibility to Pseudomonas aeruginosa lung infection in mice.BMC Genomics. 2016 May 11;17:351. doi: 10.1186/s12864-016-2676-4. BMC Genomics. 2016. PMID: 27169516 Free PMC article.
References
-
- Bragonzi A, Paroni M, Nonis A, Cramer N, Montanari S, et al. Pseudomonas aeruginosa microevolution during cystic fibrosis lung infection establishes clones with adapted virulence. AJRCCM. 2009;180:138–145. - PubMed
-
- Tümmler B. Clonal variations in Pseudomonas aeruginosa. In: Ramos J-L, Levesque RC, editors. Pseudomonas: molecular biology of emerging issues, vol. 4. New York: Springer; 2006. pp. 35–68.
-
- Bragonzi A, Wiehlmann L, Klockgether J, Cramer N, Worlitzsch D, et al. Sequence diversity of the mucABD locus in Pseudomonas aeruginosa isolates from patients with cystic fibrosis. Microbiology. 2006;152:3261–3269. - PubMed

