Gene expression noise in spatial patterning: hunchback promoter structure affects noise amplitude and distribution in Drosophila segmentation
- PMID: 21304932
- PMCID: PMC3033364
- DOI: 10.1371/journal.pcbi.1001069
Gene expression noise in spatial patterning: hunchback promoter structure affects noise amplitude and distribution in Drosophila segmentation
Abstract
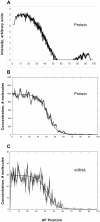
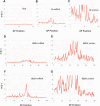
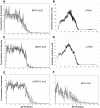
Positional information in developing embryos is specified by spatial gradients of transcriptional regulators. One of the classic systems for studying this is the activation of the hunchback (hb) gene in early fruit fly (Drosophila) segmentation by the maternally-derived gradient of the Bicoid (Bcd) protein. Gene regulation is subject to intrinsic noise which can produce variable expression. This variability must be constrained in the highly reproducible and coordinated events of development. We identify means by which noise is controlled during gene expression by characterizing the dependence of hb mRNA and protein output noise on hb promoter structure and transcriptional dynamics. We use a stochastic model of the hb promoter in which the number and strength of Bcd and Hb (self-regulatory) binding sites can be varied. Model parameters are fit to data from WT embryos, the self-regulation mutant hb(14F), and lacZ reporter constructs using different portions of the hb promoter. We have corroborated model noise predictions experimentally. The results indicate that WT (self-regulatory) Hb output noise is predominantly dependent on the transcription and translation dynamics of its own expression, rather than on Bcd fluctuations. The constructs and mutant, which lack self-regulation, indicate that the multiple Bcd binding sites in the hb promoter (and their strengths) also play a role in buffering noise. The model is robust to the variation in Bcd binding site number across a number of fly species. This study identifies particular ways in which promoter structure and regulatory dynamics reduce hb output noise. Insofar as many of these are common features of genes (e.g. multiple regulatory sites, cooperativity, self-feedback), the current results contribute to the general understanding of the reproducibility and determinacy of spatial patterning in early development.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures





Similar articles
-
Spatial bistability generates hunchback expression sharpness in the Drosophila embryo.PLoS Comput Biol. 2008 Sep 26;4(9):e1000184. doi: 10.1371/journal.pcbi.1000184. PLoS Comput Biol. 2008. PMID: 18818726 Free PMC article.
-
Live imaging of bicoid-dependent transcription in Drosophila embryos.Curr Biol. 2013 Nov 4;23(21):2135-9. doi: 10.1016/j.cub.2013.08.053. Epub 2013 Oct 17. Curr Biol. 2013. PMID: 24139736
-
3 minutes to precisely measure morphogen concentration.PLoS Genet. 2018 Oct 26;14(10):e1007676. doi: 10.1371/journal.pgen.1007676. eCollection 2018 Oct. PLoS Genet. 2018. PMID: 30365533 Free PMC article.
-
A matter of time: Formation and interpretation of the Bicoid morphogen gradient.Curr Top Dev Biol. 2020;137:79-117. doi: 10.1016/bs.ctdb.2019.11.016. Epub 2019 Dec 27. Curr Top Dev Biol. 2020. PMID: 32143754 Review.
-
Constraints and limitations on the transcriptional response downstream of the Bicoid morphogen gradient.Curr Top Dev Biol. 2020;137:119-142. doi: 10.1016/bs.ctdb.2019.12.002. Epub 2020 Jan 17. Curr Top Dev Biol. 2020. PMID: 32143741 Review.
Cited by
-
Mid-embryo patterning and precision in Drosophila segmentation: Krüppel dual regulation of hunchback.PLoS One. 2015 Mar 20;10(3):e0118450. doi: 10.1371/journal.pone.0118450. eCollection 2015. PLoS One. 2015. PMID: 25793381 Free PMC article.
-
Mean-Independent Noise Control of Cell Fates via Intermediate States.iScience. 2018 May 25;3:11-20. doi: 10.1016/j.isci.2018.04.002. Epub 2018 Apr 11. iScience. 2018. PMID: 30428314 Free PMC article.
-
Control of Hox transcription factor concentration and cell-to-cell variability by an auto-regulatory switch.Development. 2019 Jan 25;146(12):dev168179. doi: 10.1242/dev.168179. Development. 2019. PMID: 30642837 Free PMC article.
-
In silico evolution of the hunchback gene indicates redundancy in cis-regulatory organization and spatial gene expression.J Bioinform Comput Biol. 2014 Apr;12(2):1441009. doi: 10.1142/S0219720014410091. Epub 2014 Mar 25. J Bioinform Comput Biol. 2014. PMID: 24712536 Free PMC article.
-
How cells know where they are.Science. 2013 Feb 22;339(6122):923-7. doi: 10.1126/science.1224186. Science. 2013. PMID: 23430648 Free PMC article. Review.
References
-
- Houchmandzadeh B, Wieschaus E, Leibler S. Establishment of developmental precision and proportions in the early Drosophila embryo. Nature. 2002;415:798–802. - PubMed
-
- Crauk O, Dostatni N. Bicoid determines sharp and precise target gene expression on the Drosophila embryo. Curr Biol. 2005;15:1888–1898. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

