Adeno-associated viral vectors for mapping, monitoring, and manipulating neural circuits
- PMID: 21319997
- PMCID: PMC3107581
- DOI: 10.1089/hum.2010.204
Adeno-associated viral vectors for mapping, monitoring, and manipulating neural circuits
Abstract
Understanding the structure and function of neural circuits is central is neuroscience research. To address the associated questions, new genetically encoded tools have been developed for mapping, monitoring, and manipulating neurons. Essential to implementation of these tools is their selective delivery to defined neuronal populations in the brain. This has been facilitated by recent improvements in cell type-specific transgene expression using recombinant adeno-associated viral vectors. Here, we highlight these developments and discuss areas for improvement that could further expand capabilities for neural circuit analysis.
Figures


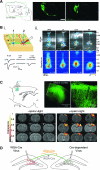
Similar articles
-
Strategies for targeting primate neural circuits with viral vectors.J Neurophysiol. 2016 Jul 1;116(1):122-34. doi: 10.1152/jn.00087.2016. Epub 2016 Apr 6. J Neurophysiol. 2016. PMID: 27052579 Free PMC article. Review.
-
Local and retrograde gene transfer into primate neuronal pathways via adeno-associated virus serotype 8 and 9.Neuroscience. 2011 Oct 13;193:249-58. doi: 10.1016/j.neuroscience.2011.06.080. Epub 2011 Jul 18. Neuroscience. 2011. PMID: 21782903
-
Translational PET applications for brain circuit mapping with transgenic neuromodulation tools.Pharmacol Biochem Behav. 2021 May;204:173147. doi: 10.1016/j.pbb.2021.173147. Epub 2021 Feb 4. Pharmacol Biochem Behav. 2021. PMID: 33549570 Free PMC article. Review.
-
Manipulating gene expression in projection-specific neuronal populations using combinatorial viral approaches.Curr Protoc Neurosci. 2013;65(435):4.35.1-20. doi: 10.1002/0471142301.ns0435s65. Curr Protoc Neurosci. 2013. PMID: 25429312 Free PMC article.
-
Use of Adeno-Associated and Herpes Simplex Viral Vectors for In Vivo Neuronal Expression in Mice.Curr Protoc Neurosci. 2015 Oct 1;73:4.37.1-4.37.31. doi: 10.1002/0471142301.ns0437s73. Curr Protoc Neurosci. 2015. PMID: 26426386 Free PMC article.
Cited by
-
Programmable Assembly of Adeno-Associated Virus-Antibody Composites for Receptor-Mediated Gene Delivery.Bioconjug Chem. 2020 Apr 15;31(4):1093-1106. doi: 10.1021/acs.bioconjchem.9b00790. Epub 2019 Dec 20. Bioconjug Chem. 2020. PMID: 31809024 Free PMC article.
-
Off-Target Expression of Cre-Dependent Adeno-Associated Viruses in Wild-Type C57BL/6J Mice.eNeuro. 2021 Nov 24;8(6):ENEURO.0363-21.2021. doi: 10.1523/ENEURO.0363-21.2021. Print 2021 Nov-Dec. eNeuro. 2021. PMID: 34785571 Free PMC article.
-
Cre-dependent selection yields AAV variants for widespread gene transfer to the adult brain.Nat Biotechnol. 2016 Feb;34(2):204-9. doi: 10.1038/nbt.3440. Epub 2016 Feb 1. Nat Biotechnol. 2016. PMID: 26829320 Free PMC article.
-
Neuromodulation in Chemosensory Pathways.Chem Senses. 2017 Jun 1;42(5):375-379. doi: 10.1093/chemse/bjx014. Chem Senses. 2017. PMID: 28379355 Free PMC article. Review.
-
CREATEd viruses go global.Nat Neurosci. 2017 Jul 26;20(8):1041-1042. doi: 10.1038/nn.4600. Nat Neurosci. 2017. PMID: 28745726 No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
