A statistical framework for modeling gene expression using chromatin features and application to modENCODE datasets
- PMID: 21324173
- PMCID: PMC3188797
- DOI: 10.1186/gb-2011-12-2-r15
A statistical framework for modeling gene expression using chromatin features and application to modENCODE datasets
Abstract
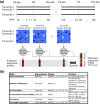
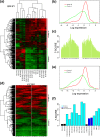
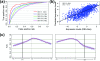
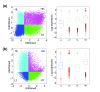
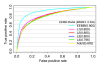
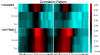
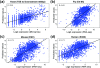
We develop a statistical framework to study the relationship between chromatin features and gene expression. This can be used to predict gene expression of protein coding genes, as well as microRNAs. We demonstrate the prediction in a variety of contexts, focusing particularly on the modENCODE worm datasets. Moreover, our framework reveals the positional contribution around genes (upstream or downstream) of distinct chromatin features to the overall prediction of expression levels.
Figures

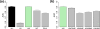

References
-
- Birney E, Stamatoyannopoulos JA, Dutta A, Guigo R, Gingeras TR, Margulies EH, Weng Z, Snyder M, Dermitzakis ET, Thurman RE, Kuehn MS, Taylor CM, Neph S, Koch CM, Asthana S, Malhotra A, Adzhubei I, Greenbaum JA, Andrews RM, Flicek P, Boyle PJ, Cao H, Carter NP, Clelland GK, Davis S, Day N, Dhami P, Dillon SC, Dorschner MO, Fiegler H. et al.Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature. 2007;447:799–816. doi: 10.1038/nature05874. - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

