Distinct properties of the XY pseudoautosomal region crucial for male meiosis
- PMID: 21330546
- PMCID: PMC3151169
- DOI: 10.1126/science.1195774
Distinct properties of the XY pseudoautosomal region crucial for male meiosis
Abstract
Meiosis requires that each chromosome find its homologous partner and undergo at least one crossover. X-Y chromosome segregation hinges on efficient crossing-over in a very small region of homology, the pseudoautosomal region (PAR). We find that mouse PAR DNA occupies unusually long chromosome axes, potentially as shorter chromatin loops, predicted to promote double-strand break (DSB) formation. Most PARs show delayed appearance of RAD51/DMC1 foci, which mark DSB ends, and all PARs undergo delayed DSB-mediated homologous pairing. Analysis of Spo11β isoform-specific transgenic mice revealed that late RAD51/DMC1 foci in the PAR are genetically distinct from both early PAR foci and global foci and that late PAR foci promote efficient X-Y pairing, recombination, and male fertility. Our findings uncover specific mechanisms that surmount the unique challenges of X-Y recombination.
Figures


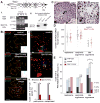
Comment in
-
Molecular biology. Hitting a tiny target in the dark.Science. 2011 Feb 18;331(6019):870-1. doi: 10.1126/science.1202373. Science. 2011. PMID: 21330526 No abstract available.
Similar articles
-
Molecular biology. Hitting a tiny target in the dark.Science. 2011 Feb 18;331(6019):870-1. doi: 10.1126/science.1202373. Science. 2011. PMID: 21330526 No abstract available.
-
Repair of Meiotic DNA Breaks and Homolog Pairing in Mouse Meiosis Requires a Minichromosome Maintenance (MCM) Paralog.Genetics. 2017 Feb;205(2):529-537. doi: 10.1534/genetics.116.196808. Epub 2016 Dec 16. Genetics. 2017. PMID: 27986806 Free PMC article.
-
Meiotic crossover control by concerted action of Rad51-Dmc1 in homolog template bias and robust homeostatic regulation.PLoS Genet. 2013;9(12):e1003978. doi: 10.1371/journal.pgen.1003978. Epub 2013 Dec 19. PLoS Genet. 2013. PMID: 24367271 Free PMC article.
-
The tricky path to recombining X and Y chromosomes in meiosis.Ann N Y Acad Sci. 2012 Sep;1267:18-23. doi: 10.1111/j.1749-6632.2012.06593.x. Ann N Y Acad Sci. 2012. PMID: 22954211 Free PMC article. Review.
-
Recent advances in mechanisms ensuring the pairing, synapsis and segregation of XY chromosomes in mice and humans.Cell Mol Life Sci. 2024 Apr 23;81(1):194. doi: 10.1007/s00018-024-05216-0. Cell Mol Life Sci. 2024. PMID: 38653846 Free PMC article. Review.
Cited by
-
Mechanistic basis of infertility of mouse intersubspecific hybrids.Proc Natl Acad Sci U S A. 2013 Feb 5;110(6):E468-77. doi: 10.1073/pnas.1219126110. Epub 2013 Jan 17. Proc Natl Acad Sci U S A. 2013. PMID: 23329330 Free PMC article.
-
Gradual implementation of the meiotic recombination program via checkpoint pathways controlled by global DSB levels.Mol Cell. 2015 Mar 5;57(5):797-811. doi: 10.1016/j.molcel.2014.12.027. Epub 2015 Feb 5. Mol Cell. 2015. PMID: 25661491 Free PMC article.
-
The Y chromosome of the Okinawa spiny rat, Tokudaia muenninki, was rescued through fusion with an autosome.Chromosome Res. 2012 Jan;20(1):111-25. doi: 10.1007/s10577-011-9268-6. Chromosome Res. 2012. PMID: 22198613
-
Signatures of replication timing, recombination, and sex in the spectrum of rare variants on the human X chromosome and autosomes.Proc Natl Acad Sci U S A. 2019 Sep 3;116(36):17916-17924. doi: 10.1073/pnas.1900714116. Epub 2019 Aug 19. Proc Natl Acad Sci U S A. 2019. PMID: 31427530 Free PMC article.
-
Cyclin B3 is dispensable for mouse spermatogenesis.Chromosoma. 2019 Sep;128(3):473-487. doi: 10.1007/s00412-019-00725-5. Epub 2019 Aug 24. Chromosoma. 2019. PMID: 31446450 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

