The Gα subunit signals through the Ste50 protein during the mating pheromone response in the yeast Kluyveromyces lactis
- PMID: 21335532
- PMCID: PMC3127637
- DOI: 10.1128/EC.00285-10
The Gα subunit signals through the Ste50 protein during the mating pheromone response in the yeast Kluyveromyces lactis
Abstract
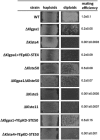
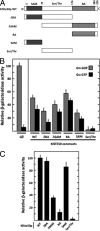
Yeast mating signal transduction pathways require a heterotrimeric G protein composed of Gα, Gβ, and Gγ subunits connected to a mitogen-activated protein kinase (MAPK) module. While in Saccharomyces cerevisiae elimination of Gα induces constitutive activation of the mating pathway, in Kluyveromyces lactis it produces partial sterility, which indicates that K. lactis Gα (KlGα) is required to positively activate mating. We use physical interaction experiments to determine that KlGα interacts with the adaptor protein KlSte50p. The Ras association (RA) domain of KlSte50p favored interaction with the GDP-bound KlGα subunit, and when the KlGα protein is constitutively activated, the interaction drops significantly. Additionally, KlSte50p strongly associates with the MAPK kinase kinase (MAPKKK) KlSte11p through its sterile alpha motif (SAM) domain. Genetic experiments placed KlSte50p downstream of the G protein α subunit, indicating that KlGα may stimulate the mating pathway via KlSte50p. Fusion of KlSte50p to the KlGβ subunit partially eliminated the requirement of KlGα for mating, indicating that one contribution of KlGα to the mating pathway is to facilitate plasma membrane anchoring of KlSte50p.
Figures


Similar articles
-
The beta subunit of the heterotrimeric G protein triggers the Kluyveromyces lactis pheromone response pathway in the absence of the gamma subunit.Mol Biol Cell. 2010 Feb 1;21(3):489-98. doi: 10.1091/mbc.e09-06-0472. Epub 2009 Dec 16. Mol Biol Cell. 2010. PMID: 20016006 Free PMC article.
-
The Gbeta(KlSte4p) subunit of the heterotrimeric G protein has a positive and essential role in the induction of mating in the yeast Kluyveromyces lactis.Yeast. 2005 Sep;22(12):947-56. doi: 10.1002/yea.1278. Yeast. 2005. PMID: 16134098
-
Protein kinases involved in mating and osmotic stress in the yeast Kluyveromyces lactis.Eukaryot Cell. 2008 Jan;7(1):78-85. doi: 10.1128/EC.00362-07. Epub 2007 Nov 16. Eukaryot Cell. 2008. PMID: 18024598 Free PMC article.
-
The pheromone response pathway of Kluyveromyces lactis.FEMS Yeast Res. 2006 May;6(3):336-44. doi: 10.1111/j.1567-1364.2005.00022.x. FEMS Yeast Res. 2006. PMID: 16630274 Review.
-
The role of adaptor protein Ste50-dependent regulation of the MAPKKK Ste11 in multiple signalling pathways of yeast.Curr Genet. 2003 Jun;43(3):161-70. doi: 10.1007/s00294-003-0383-6. Epub 2003 Mar 11. Curr Genet. 2003. PMID: 12764668 Review.
Cited by
-
Mate and fuse: how yeast cells do it.Open Biol. 2013 Mar 6;3(3):130008. doi: 10.1098/rsob.130008. Open Biol. 2013. PMID: 23466674 Free PMC article. Review.
References
-
- Bardwell L. 2004. A walk-through of the yeast mating pheromone response pathway. Peptides 25:1465–1476 - PubMed
-
- Bennett R. J., Johnson A. D. 2006. The role of nutrient regulation and the Gpa2 protein in the mating pheromone response of C. albicans. Mol. Microbiol. 62:100–119 - PubMed
-
- Blinder D., Bouvier S., Jenness D. D. 1989. Constitutive mutants in the yeast pheromone response: ordered function of the gene products. Cell 56:479–486 - PubMed
-
- Dietzel C., Kurjan J. 1987. The yeast Scg1 gene: a Gα-like protein implicated in the a- and α-factor response pathway. Cell 50:1001–1010 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

