Hybrid metabolic flux analysis: combining stoichiometric and statistical constraints to model the formation of complex recombinant products
- PMID: 21352531
- PMCID: PMC3236310
- DOI: 10.1186/1752-0509-5-34
Hybrid metabolic flux analysis: combining stoichiometric and statistical constraints to model the formation of complex recombinant products
Abstract
Background: Stoichiometric models constitute the basic framework for fluxome quantification in the realm of metabolic engineering. A recurrent bottleneck, however, is the establishment of consistent stoichiometric models for the synthesis of recombinant proteins or viruses. Although optimization algorithms for in silico metabolic redesign have been developed in the context of genome-scale stoichiometric models for small molecule production, still rudimentary knowledge of how different cellular levels are regulated and phenotypically expressed prevents their full applicability for complex product optimization.
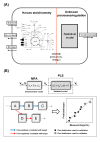
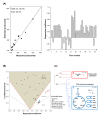
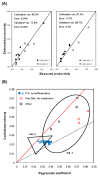
Results: A hybrid framework is presented combining classical metabolic flux analysis with projection to latent structures to further link estimated metabolic fluxes with measured productivities. We first explore the functional metabolic decomposition of a baculovirus-producing insect cell line from experimental data, highlighting the TCA cycle and mitochondrial respiration as pathways strongly associated with viral replication. To reduce uncertainty in metabolic target identification, a Monte Carlo sampling method was used to select meaningful associations with the target, from which 66% of the estimated fluxome had to be screened out due to weak correlations and/or high estimation errors. The proposed hybrid model was then validated using a subset of preliminary experiments to pinpoint the same determinant pathways, while predicting the productivity of independent cultures.
Conclusions: Overall, the results indicate our hybrid metabolic flux analysis framework is an advantageous tool for metabolic identification and quantification in incomplete or ill-defined metabolic networks. As experimental and computational solutions for constructing comprehensive global cellular models are in development, the contribution of hybrid metabolic flux analysis should constitute a valuable complement to current computational platforms in bridging the metabolic state with improved cell culture performance.
Figures
Similar articles
-
The role of host cell physiology in the productivity of the baculovirus-insect cell system: Fluxome analysis of Trichoplusia ni and Spodoptera frugiperda cell lines.Biotechnol Bioeng. 2017 Mar;114(3):674-684. doi: 10.1002/bit.26089. Epub 2016 Sep 26. Biotechnol Bioeng. 2017. PMID: 27568545
-
A hybrid model of anaerobic E. coli GJT001: combination of elementary flux modes and cybernetic variables.Biotechnol Prog. 2008 Sep-Oct;24(5):993-1006. doi: 10.1002/btpr.73. Biotechnol Prog. 2008. PMID: 19194908
-
Hybrid elementary flux analysis/nonparametric modeling: application for bioprocess control.BMC Bioinformatics. 2007 Jan 29;8:30. doi: 10.1186/1471-2105-8-30. BMC Bioinformatics. 2007. PMID: 17261182 Free PMC article.
-
Metabolic flux analysis and metabolic engineering of microorganisms.Mol Biosyst. 2008 Feb;4(2):113-20. doi: 10.1039/b712395g. Epub 2007 Oct 16. Mol Biosyst. 2008. PMID: 18213404 Review.
-
Advanced stoichiometric analysis of metabolic networks of mammalian systems.Crit Rev Biomed Eng. 2011;39(6):511-34. doi: 10.1615/critrevbiomedeng.v39.i6.30. Crit Rev Biomed Eng. 2011. PMID: 22196224 Free PMC article. Review.
Cited by
-
Towards a widespread adoption of metabolic modeling tools in biopharmaceutical industry: a process systems biology engineering perspective.NPJ Syst Biol Appl. 2020 Mar 13;6(1):6. doi: 10.1038/s41540-020-0127-y. NPJ Syst Biol Appl. 2020. PMID: 32170148 Free PMC article. Review.
-
From Spatial-Temporal Multiscale Modeling to Application: Bridging the Valley of Death in Industrial Biotechnology.Bioengineering (Basel). 2023 Jun 20;10(6):744. doi: 10.3390/bioengineering10060744. Bioengineering (Basel). 2023. PMID: 37370675 Free PMC article. Review.
-
Metabolic Flux Analysis-Linking Isotope Labeling and Metabolic Fluxes.Metabolites. 2020 Nov 6;10(11):447. doi: 10.3390/metabo10110447. Metabolites. 2020. PMID: 33172051 Free PMC article. Review.
-
Toward system-level understanding of baculovirus-host cell interactions: from molecular fundamental studies to large-scale proteomics approaches.Front Microbiol. 2012 Nov 9;3:391. doi: 10.3389/fmicb.2012.00391. eCollection 2012. Front Microbiol. 2012. PMID: 23162544 Free PMC article.
-
Hybrid semiparametric systems for quantitative sequence-activity modeling of synthetic biological parts.Synth Biol (Oxf). 2018 Jun 26;3(1):ysy010. doi: 10.1093/synbio/ysy010. eCollection 2018. Synth Biol (Oxf). 2018. PMID: 32995518 Free PMC article.
References
-
- Lee SY, Hong SH, Lee DY, Kim TY. In: Bioinformatics Technologies. Yi-Ping Phoebe Chen, editor. Springer Berlin Heidelberg; 2005. Systems biotechnology: a new paradigm in biotechnology development; pp. 155–177. full_text.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

