Change is good: variations in common biological mechanisms in the epsilonproteobacterial genera Campylobacter and Helicobacter
- PMID: 21372321
- PMCID: PMC3063351
- DOI: 10.1128/MMBR.00035-10
Change is good: variations in common biological mechanisms in the epsilonproteobacterial genera Campylobacter and Helicobacter
Abstract
Microbial evolution and subsequent species diversification enable bacterial organisms to perform common biological processes by a variety of means. The epsilonproteobacteria are a diverse class of prokaryotes that thrive in diverse habitats. Many of these environmental niches are labeled as extreme, whereas other niches include various sites within human, animal, and insect hosts. Some epsilonproteobacteria, such as Campylobacter jejuni and Helicobacter pylori, are common pathogens of humans that inhabit specific regions of the gastrointestinal tract. As such, the biological processes of pathogenic Campylobacter and Helicobacter spp. are often modeled after those of common enteric pathogens such as Salmonella spp. and Escherichia coli. While many exquisite biological mechanisms involving biochemical processes, genetic regulatory pathways, and pathogenesis of disease have been elucidated from studies of Salmonella spp. and E. coli, these paradigms often do not apply to the same processes in the epsilonproteobacteria. Instead, these bacteria often display extensive variation in common biological mechanisms relative to those of other prototypical bacteria. In this review, five biological processes of commonly studied model bacterial species are compared to those of the epsilonproteobacteria C. jejuni and H. pylori. Distinct differences in the processes of flagellar biosynthesis, DNA uptake and recombination, iron homeostasis, interaction with epithelial cells, and protein glycosylation are highlighted. Collectively, these studies support a broader view of the vast repertoire of biological mechanisms employed by bacteria and suggest that future studies of the epsilonproteobacteria will continue to provide novel and interesting information regarding prokaryotic cellular biology.
Figures




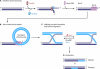

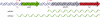




References
-
- Aas, F. E., et al. 2002. Competence for natural transformation in Neisseria gonorrhoeae: components of DNA binding and uptake linked to type IV pilus expression. Mol. Microbiol. 46:749-760. - PubMed
-
- Abrahams, G. L., P. Muller, and M. Hensel. 2006. Functional dissection of SseF, a type III effector protein involved in positioning the Salmonella-containing vacuole. Traffic 7:950-965. - PubMed
-
- Abu-Qarn, M., J. Eichler, and N. Sharon. 2008. Not just for Eukarya anymore: protein glycosylation in Bacteria and Archaea. Curr. Opin. Struct. Biol. 18:544-550. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

