The organellar genome and metabolic potential of the hydrogen-producing mitochondrion of Nyctotherus ovalis
- PMID: 21378103
- PMCID: PMC3144386
- DOI: 10.1093/molbev/msr059
The organellar genome and metabolic potential of the hydrogen-producing mitochondrion of Nyctotherus ovalis
Abstract
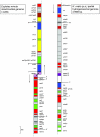
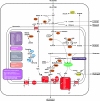
It is generally accepted that hydrogenosomes (hydrogen-producing organelles) evolved from a mitochondrial ancestor. However, until recently, only indirect evidence for this hypothesis was available. Here, we present the almost complete genome of the hydrogen-producing mitochondrion of the anaerobic ciliate Nyctotherus ovalis and show that, except for the notable absence of genes encoding electron transport chain components of Complexes III, IV, and V, it has a gene content similar to the mitochondrial genomes of aerobic ciliates. Analysis of the genome of the hydrogen-producing mitochondrion, in combination with that of more than 9,000 genomic DNA and cDNA sequences, allows a preliminary reconstruction of the organellar metabolism. The sequence data indicate that N. ovalis possesses hydrogen-producing mitochondria that have a truncated, two step (Complex I and II) electron transport chain that uses fumarate as electron acceptor. In addition, components of an extensive protein network for the metabolism of amino acids, defense against oxidative stress, mitochondrial protein synthesis, mitochondrial protein import and processing, and transport of metabolites across the mitochondrial membrane were identified. Genes for MPV17 and ACN9, two hypothetical proteins linked to mitochondrial disease in humans, were also found. The inferred metabolism is remarkably similar to the organellar metabolism of the phylogenetically distant anaerobic Stramenopile Blastocystis. Notably, the Blastocystis organelle and that of the related flagellate Proteromonas lacertae also lack genes encoding components of Complexes III, IV, and V. Thus, our data show that the hydrogenosomes of N. ovalis are highly specialized hydrogen-producing mitochondria.
Figures

Similar articles
-
An anaerobic mitochondrion that produces hydrogen.Nature. 2005 Mar 3;434(7029):74-9. doi: 10.1038/nature03343. Nature. 2005. PMID: 15744302
-
Organelles in Blastocystis that blur the distinction between mitochondria and hydrogenosomes.Curr Biol. 2008 Apr 22;18(8):580-5. doi: 10.1016/j.cub.2008.03.037. Epub 2008 Apr 10. Curr Biol. 2008. PMID: 18403202 Free PMC article.
-
Analysis of two genomes from the mitochondrion-like organelle of the intestinal parasite Blastocystis: complete sequences, gene content, and genome organization.Mol Biol Evol. 2008 Nov;25(11):2475-82. doi: 10.1093/molbev/msn193. Epub 2008 Sep 2. Mol Biol Evol. 2008. PMID: 18765437 Free PMC article.
-
Diversity and reductive evolution of mitochondria among microbial eukaryotes.Philos Trans R Soc Lond B Biol Sci. 2010 Mar 12;365(1541):713-27. doi: 10.1098/rstb.2009.0224. Philos Trans R Soc Lond B Biol Sci. 2010. PMID: 20124340 Free PMC article. Review.
-
Hydrogenosomes under microscopy.Tissue Cell. 2009 Jun;41(3):151-68. doi: 10.1016/j.tice.2009.01.001. Epub 2009 Mar 17. Tissue Cell. 2009. PMID: 19297000 Review.
Cited by
-
Diversity and origins of anaerobic metabolism in mitochondria and related organelles.Philos Trans R Soc Lond B Biol Sci. 2015 Sep 26;370(1678):20140326. doi: 10.1098/rstb.2014.0326. Philos Trans R Soc Lond B Biol Sci. 2015. PMID: 26323757 Free PMC article. Review.
-
The Oxytricha trifallax mitochondrial genome.Genome Biol Evol. 2012;4(2):136-54. doi: 10.1093/gbe/evr136. Epub 2011 Dec 16. Genome Biol Evol. 2012. PMID: 22179582 Free PMC article.
-
Population Genetics of Paramecium Mitochondrial Genomes: Recombination, Mutation Spectrum, and Efficacy of Selection.Genome Biol Evol. 2019 May 1;11(5):1398-1416. doi: 10.1093/gbe/evz081. Genome Biol Evol. 2019. PMID: 30980669 Free PMC article.
-
The evolution of early cellular systems viewed through the lens of biological interactions.Front Microbiol. 2015 Oct 19;6:1144. doi: 10.3389/fmicb.2015.01144. eCollection 2015. Front Microbiol. 2015. PMID: 26539175 Free PMC article.
-
Mitogenomic architecture and evolution of the soil ciliates Colpoda.mSystems. 2024 Feb 20;9(2):e0116123. doi: 10.1128/msystems.01161-23. Epub 2024 Jan 23. mSystems. 2024. PMID: 38259100 Free PMC article.
References
-
- Abascal F, Zardoya R, Posada D. ProtTest: selection of best-fit models of protein evolution. Bioinformatics. 2005;21:2104–2105. - PubMed
-
- Aguilera P, Barry T, Tovar J. Entamoeba histolytica mitosomes: organelles in search of a function. Exp Parasitol. 2008;118:10–16. - PubMed
-
- Akhmanova A, Voncken FG, Harhangi H, Hosea KM, Vogels GD, Hackstein JH. Cytosolic enzymes with a mitochondrial ancestry from the anaerobic chytrid Piromyces sp. E2. Mol Microbiol. 1998;30:1017–1027. - PubMed
-
- Akhmanova A, Voncken FG, Hosea KM, Harhangi H, Keltjens JT, op den Camp HJ, Vogels GD, Hackstein JH. A hydrogenosome with pyruvate formate-lyase: anaerobic chytrid fungi use an alternative route for pyruvate catabolism. Mol Microbiol. 1999;32:1103–1114. - PubMed
-
- Akhmanova A, Voncken F, van Alen T, van Hoek A, Boxma B, Vogels G, Veenhuis M, Hackstein JH. A hydrogenosome with a genome. Nature. 1998;396:527–528. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

