Analysis of molecular diffusion by first-passage time variance identifies the size of confinement zones
- PMID: 21402028
- PMCID: PMC3059732
- DOI: 10.1016/j.bpj.2011.01.064
Analysis of molecular diffusion by first-passage time variance identifies the size of confinement zones
Abstract
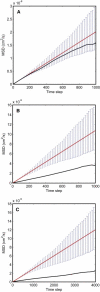
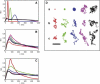
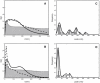
The diffusion of receptors within the two-dimensional environment of the plasma membrane is a complex process. Although certain components diffuse according to a random walk model (Brownian diffusion), an overwhelming body of work has found that membrane diffusion is nonideal (anomalous diffusion). One of the most powerful methods for studying membrane diffusion is single particle tracking (SPT), which records the trajectory of a label attached to a membrane component of interest. One of the outstanding problems in SPT is the analysis of data to identify the presence of heterogeneity. We have adapted a first-passage time (FPT) algorithm, originally developed for the interpretation of animal movement, for the analysis of SPT data. We discuss the general application of the FPT analysis to molecular diffusion, and use simulations to test the method against data containing known regions of confinement. We conclude that FPT can be used to identify the presence and size of confinement within trajectories of the receptor LFA-1, and these results are consistent with previous reports on the size of LFA-1 clusters. The analysis of trajectory data for cell surface receptors by FPT provides a robust method to determine the presence and size of confined regions of diffusion.
Copyright © 2011 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures

Similar articles
-
Time series analysis of particle tracking data for molecular motion on the cell membrane.Bull Math Biol. 2009 Nov;71(8):1967-2024. doi: 10.1007/s11538-009-9434-6. Epub 2009 Aug 6. Bull Math Biol. 2009. PMID: 19657701 Free PMC article.
-
Automatic detection of diffusion modes within biological membranes using back-propagation neural network.BMC Bioinformatics. 2016 May 4;17(1):197. doi: 10.1186/s12859-016-1064-z. BMC Bioinformatics. 2016. PMID: 27141816 Free PMC article.
-
Confined lateral diffusion of membrane receptors as studied by single particle tracking (nanovid microscopy). Effects of calcium-induced differentiation in cultured epithelial cells.Biophys J. 1993 Nov;65(5):2021-40. doi: 10.1016/S0006-3495(93)81253-0. Biophys J. 1993. PMID: 8298032 Free PMC article.
-
Interprotein interactions are responsible for the confined diffusion of a G-protein-coupled receptor at the cell surface.Biochem Soc Trans. 2003 Oct;31(Pt 5):1001-5. doi: 10.1042/bst0311001. Biochem Soc Trans. 2003. PMID: 14505468 Review.
-
Mechanisms underlying anomalous diffusion in the plasma membrane.Curr Top Membr. 2015;75:167-207. doi: 10.1016/bs.ctm.2015.03.002. Epub 2015 Apr 15. Curr Top Membr. 2015. PMID: 26015283 Review.
Cited by
-
Precise Detection and Visualization of Nanoscale Temporal Confinement in Single-Molecule Tracking Analysis.Membranes (Basel). 2022 Jun 24;12(7):650. doi: 10.3390/membranes12070650. Membranes (Basel). 2022. PMID: 35877853 Free PMC article.
-
A Simple and Powerful Analysis of Lateral Subdiffusion Using Single Particle Tracking.Biophys J. 2017 Dec 5;113(11):2452-2463. doi: 10.1016/j.bpj.2017.09.017. Biophys J. 2017. PMID: 29211999 Free PMC article.
-
Simulation of receptor triggering by kinetic segregation shows role of oligomers and close contacts.Biophys J. 2022 May 3;121(9):1660-1674. doi: 10.1016/j.bpj.2022.03.033. Epub 2022 Mar 31. Biophys J. 2022. PMID: 35367423 Free PMC article.
-
Detection of Diffusion Heterogeneity in Single Particle Tracking Trajectories Using a Hidden Markov Model with Measurement Noise Propagation.PLoS One. 2015 Oct 16;10(10):e0140759. doi: 10.1371/journal.pone.0140759. eCollection 2015. PLoS One. 2015. PMID: 26473352 Free PMC article.
-
A Hidden Markov Model for Detecting Confinement in Single-Particle Tracking Trajectories.Biophys J. 2018 Nov 6;115(9):1741-1754. doi: 10.1016/j.bpj.2018.09.005. Epub 2018 Sep 13. Biophys J. 2018. PMID: 30274829 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

