Backbone and side-chain contributions in protein denaturation by urea
- PMID: 21402035
- PMCID: PMC3059734
- DOI: 10.1016/j.bpj.2011.01.028
Backbone and side-chain contributions in protein denaturation by urea
Abstract
Urea is a commonly used protein denaturant, and it is of great interest to determine its interaction with various protein groups to elucidate the molecular basis of its effect on protein stability. Using the Trp-cage miniprotein as a model system, we report what we believe to be the first computation of changes in the preferential interaction coefficient of the protein upon urea denaturation from molecular-dynamics simulations and examine the contributions from the backbone and the side-chain groups. The preferential interaction is obtained from reversible folding/unfolding replica exchange molecular-dynamics simulations of Trp-cage in presence of urea, over a wide range of urea concentration. The increase in preferential interaction upon unfolding is dominated by the side-chain contribution, rather than the backbone. Similar trends are observed in simulations using two different force fields, Amber94 and Amber99sb, for the protein. The magnitudes of the side-chain and backbone contributions differ in the two force fields, despite containing identical protein-solvent interaction terms. The differences arise from the unfolded ensembles sampled, with Amber99sb favoring conformations with larger surface area and lower helical content. These results emphasize the importance of the side-chain interactions with urea in protein denaturation, and highlight the dependence of the computed driving forces on the unfolded ensemble sampled.
Copyright © 2011 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures


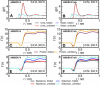



References
-
- Timasheff S.N. The control of protein stability and association by weak interactions with water: how do solvents affect these processes? Annu. Rev. Biophys. Biomol. Struct. 1993;22:67–97. - PubMed
-
- Yancey P.H., Clark M.E., Somero G.N. Living with water stress: evolution of osmolyte systems. Science. 1982;217:1214–1222. - PubMed
-
- Kauzmann W. Some factors in the interpretation of protein denaturation. Adv. Protein Chem. 1959;14:1–63. - PubMed
-
- Pace C.N. Determination and analysis of urea and guanidine hydrochloride denaturation curves. Methods Enzymol. 1986;131:266–280. - PubMed
-
- Frank H.S., Franks F. Structural approach to solvent power of water for hydrocarbons—urea as a structure breaker. J. Chem. Phys. 1968;48:4746–4757.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

