Structural changes in bacteriorhodopsin during in vitro refolding from a partially denatured state
- PMID: 21402039
- PMCID: PMC3059736
- DOI: 10.1016/j.bpj.2011.02.004
Structural changes in bacteriorhodopsin during in vitro refolding from a partially denatured state
Abstract
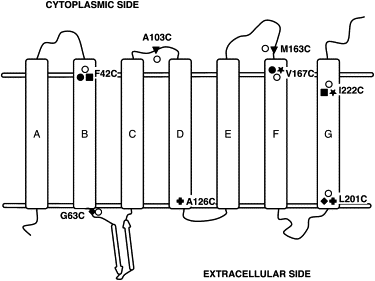
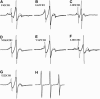
We report on the formation of the secondary and tertiary structure of bacteriorhodopsin during its in vitro refolding from an SDS-denatured state. We used the mobility of single spin labels in seven samples, attached at various locations to six of the seven helical segments to engineered cysteine residues, to follow coil-to-helix formation. Distance measurements obtained by spin dipolar quenching in six samples labeled at either the cytoplasmic or extracellular ends of pairs of helices revealed the time dependence of the recovery of the transmembrane helical bundle. The secondary structure in the majority of the helical segments refolds with a time constant of <100-140 ms. Recovery of the tertiary structure is achieved by sequential association of the helices and occurs in at least three distinct steps with time constants of 1), well below 1 s; 2), 3-4 s; and 3), 60-130 s (the latter depending on the helical pair). The slowest of these processes occurs in concert with recovery of the retinal chromophore.
Copyright © 2011 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures



Similar articles
-
Secondary and tertiary structure of bacteriorhodopsin in the SDS denatured state.Biochemistry. 2012 Feb 14;51(6):1051-60. doi: 10.1021/bi201769z. Epub 2012 Jan 30. Biochemistry. 2012. PMID: 22242919
-
Kinetics and motional dynamics of spin-labeled yeast iso-1-cytochrome c: 1. Stopped-flow electron paramagnetic resonance as a probe for protein folding/unfolding of the C-terminal helix spin-labeled at cysteine 102.Biochemistry. 1997 Mar 11;36(10):2884-97. doi: 10.1021/bi962155i. Biochemistry. 1997. PMID: 9062118
-
Characterization of Denatured States and Reversible Unfolding of Sensory Rhodopsin II.J Mol Biol. 2018 Oct 19;430(21):4068-4086. doi: 10.1016/j.jmb.2018.07.031. Epub 2018 Aug 9. J Mol Biol. 2018. PMID: 30098339
-
Unravelling the folding of bacteriorhodopsin.Biochim Biophys Acta. 2000 Aug 30;1460(1):4-14. doi: 10.1016/s0005-2728(00)00125-0. Biochim Biophys Acta. 2000. PMID: 10984586 Review.
-
Retinal proteins as model systems for membrane protein folding.Biochim Biophys Acta. 2014 May;1837(5):656-63. doi: 10.1016/j.bbabio.2013.11.021. Epub 2013 Dec 12. Biochim Biophys Acta. 2014. PMID: 24333783 Review.
Cited by
-
Lipid bilayer composition modulates the unfolding free energy of a knotted α-helical membrane protein.Proc Natl Acad Sci U S A. 2018 Feb 20;115(8):E1799-E1808. doi: 10.1073/pnas.1714668115. Epub 2018 Feb 5. Proc Natl Acad Sci U S A. 2018. PMID: 29432185 Free PMC article.
-
How physical forces drive the process of helical membrane protein folding.EMBO Rep. 2022 Feb 3;23(3):e53025. doi: 10.15252/embr.202153025. Epub 2022 Feb 8. EMBO Rep. 2022. PMID: 35133709 Free PMC article. Review.
-
The safety dance: biophysics of membrane protein folding and misfolding in a cellular context.Q Rev Biophys. 2015 Feb;48(1):1-34. doi: 10.1017/S0033583514000110. Epub 2014 Nov 25. Q Rev Biophys. 2015. PMID: 25420508 Free PMC article.
-
Unpaired electrons as probes of catalytic systems.Sci Prog. 2014;97(Pt 4):303-70. doi: 10.3184/003685014X14151169734173. Sci Prog. 2014. PMID: 25642491 Free PMC article.
-
Bacteriorhodopsin folds through a poorly organized transition state.J Am Chem Soc. 2014 Nov 26;136(47):16574-81. doi: 10.1021/ja508359n. Epub 2014 Nov 17. J Am Chem Soc. 2014. PMID: 25369295 Free PMC article.
References
-
- Huang K.S., Bayley H., Khorana H.G. Refolding of an integral membrane protein. Denaturation, renaturation, and reconstitution of intact bacteriorhodopsin and two proteolytic fragments. J. Biol. Chem. 1981;256:3802–3809. - PubMed
-
- London E., Khorana H.G. Denaturation and renaturation of bacteriorhodopsin in detergents and lipid-detergent mixtures. J. Biol. Chem. 1982;257:7003–7011. - PubMed
-
- Liao M.J., London E., Khorana H.G. Regeneration of the native bacteriorhodopsin structure from two chymotryptic fragments. J. Biol. Chem. 1983;258:9949–9955. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

