Identification of genes involved in cell wall biogenesis in grasses by differential gene expression profiling of elongating and non-elongating maize internodes
- PMID: 21402660
- PMCID: PMC3130177
- DOI: 10.1093/jxb/err045
Identification of genes involved in cell wall biogenesis in grasses by differential gene expression profiling of elongating and non-elongating maize internodes
Abstract
Despite the economic importance of grasses as food, feed, and energy crops, little is known about the genes that control their cell wall synthesis, assembly, and remodelling. Here a detailed transcriptome analysis that allowed the identification of genes involved in grass cell wall biogenesis is provided. Differential gene expression profiling, using maize oligonucleotide arrays, was used to identify genes differentially expressed between an elongating internode, containing cells exhibiting primary cell wall synthesis, and an internode that had just ceased elongation and in which many cells were depositing secondary cell wall material. This is one of only a few studies specifically aimed at the identification of cell wall-related genes in grasses. Analysis identified new candidate genes for a role in primary and secondary cell wall biogenesis in grasses. The results suggest that many proteins involved in cell wall processes during normal development are also recruited during defence-related cell wall remodelling events. This work provides a platform for studies in which candidate genes will be functionally tested for involvement in cell wall-related processes, increasing our knowledge of cell wall biogenesis and its regulation in grasses. Since several grasses are currently being developed as lignocellulosic feedstocks for biofuel production, this improved understanding of grass cell wall biogenesis is timely, as it will facilitate the manipulation of traits favourable for sustainable food and biofuel production.
Figures


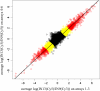

References
-
- Al-Ghazi Y, Bourot S, Arioli T, Dennis ES, Llewellyn DJ. Transcript profiling during fiber development identifies pathways in secondary metabolism and cell wall structure that may contribute to cotton fiber quality. Plant and Cell Physiology. 2009;50:1364–1381. - PubMed
-
- Andersson-Gunneras S, Mellerowicz EJ, Love J, Segerman B, Ohmiya Y, Coutinho PM, Nilsson P, Henrissat B, Moritz T, Sundberg B. Biosynthesis of cellulose-enriched tension wood in populus: global analysis of transcripts and metabolites identifies biochemical and developmental regulators in secondary wall biosynthesis. The Plant Journal. 2006;45:144–165. - PubMed
-
- Appenzeller L, Doblin M, Barreiro R, Wang HY, Niu XM, Kollipara K, Carrigan L, Tomes D, Chapman M, Dhugga KS. Cellulose synthesis in maize: isolation and expression analysis of the cellulose synthase (CesA) gene family. Cellulose. 2004;11:287–299.
-
- Benjamini Y, Hochberg Y. Controlling the false discovery rate—a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society B: Methodological. 1995;57:289–300.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

