Regulation of efflux pump expression and drug resistance by the transcription factors Mrr1, Upc2, and Cap1 in Candida albicans
- PMID: 21402859
- PMCID: PMC3088179
- DOI: 10.1128/AAC.01343-10
Regulation of efflux pump expression and drug resistance by the transcription factors Mrr1, Upc2, and Cap1 in Candida albicans
Abstract
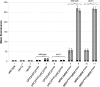
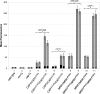
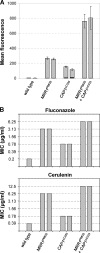
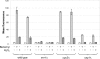
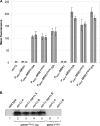
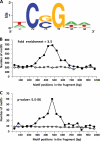
Constitutive overexpression of the Mdr1 efflux pump is an important mechanism of acquired drug resistance in the yeast Candida albicans. The zinc cluster transcription factor Mrr1 is a central regulator of MDR1 expression, but other transcription factors have also been implicated in MDR1 regulation. To better understand how MDR1-mediated drug resistance is achieved in this fungal pathogen, we studied the interdependence of Mrr1 and two other MDR1 regulators, Upc2 and Cap1, in the control of MDR1 expression. A mutated, constitutively active Mrr1 could upregulate MDR1 and confer drug resistance in the absence of Upc2 or Cap1. On the other hand, Upc2 containing a gain-of-function mutation only slightly activated the MDR1 promoter, and this activation depended on the presence of a functional MRR1 gene. In contrast, a C-terminally truncated, activated form of Cap1 could upregulate MDR1 in a partially Mrr1-independent fashion. The induction of MDR1 expression by toxic chemicals occurred independently of Upc2 but required the presence of Mrr1 and also partially depended on Cap1. Transcriptional profiling and in vivo DNA binding studies showed that a constitutively active Mrr1 binds to and upregulates most of its direct target genes in the presence or absence of Cap1. Therefore, Mrr1 and Cap1 cooperate in the environmental induction of MDR1 expression in wild-type C. albicans, but gain-of-function mutations in either of the two transcription factors can independently mediate efflux pump overexpression and drug resistance.
Figures
Similar articles
-
Differential requirement of the transcription factor Mcm1 for activation of the Candida albicans multidrug efflux pump MDR1 by its regulators Mrr1 and Cap1.Antimicrob Agents Chemother. 2011 May;55(5):2061-6. doi: 10.1128/AAC.01467-10. Epub 2011 Feb 22. Antimicrob Agents Chemother. 2011. PMID: 21343453 Free PMC article.
-
SAGA/ADA complex subunit Ada2 is required for Cap1- but not Mrr1-mediated upregulation of the Candida albicans multidrug efflux pump MDR1.Antimicrob Agents Chemother. 2014 Sep;58(9):5102-10. doi: 10.1128/AAC.03065-14. Epub 2014 Jun 16. Antimicrob Agents Chemother. 2014. PMID: 24936593 Free PMC article.
-
Mutations in the multi-drug resistance regulator MRR1, followed by loss of heterozygosity, are the main cause of MDR1 overexpression in fluconazole-resistant Candida albicans strains.Mol Microbiol. 2008 Aug;69(4):827-40. doi: 10.1111/j.1365-2958.2008.06309.x. Epub 2008 May 27. Mol Microbiol. 2008. PMID: 18577180 Free PMC article.
-
[Advances in the study of Candida albicans gene mutation on azole drug resistance].Yao Xue Xue Bao. 2010 Jul;45(7):821-6. Yao Xue Xue Bao. 2010. PMID: 20931777 Review. Chinese.
-
The genetic basis of fluconazole resistance development in Candida albicans.Biochim Biophys Acta. 2002 Jul 18;1587(2-3):240-8. doi: 10.1016/s0925-4439(02)00087-x. Biochim Biophys Acta. 2002. PMID: 12084466 Review.
Cited by
-
Molecular Mechanisms Associated with Antifungal Resistance in Pathogenic Candida Species.Cells. 2023 Nov 19;12(22):2655. doi: 10.3390/cells12222655. Cells. 2023. PMID: 37998390 Free PMC article. Review.
-
Candida parapsilosis Virulence and Antifungal Resistance Mechanisms: A Comprehensive Review of Key Determinants.J Fungi (Basel). 2023 Jan 5;9(1):80. doi: 10.3390/jof9010080. J Fungi (Basel). 2023. PMID: 36675901 Free PMC article. Review.
-
Prevalence and mechanisms of azole resistance in clinical isolates of Aspergillus section Fumigati species in a Canadian tertiary care centre, 2000 to 2013.J Antimicrob Chemother. 2020 Apr 1;75(4):849-858. doi: 10.1093/jac/dkz534. J Antimicrob Chemother. 2020. PMID: 31891387 Free PMC article.
-
Molecular and genetic basis of azole antifungal resistance in the opportunistic pathogenic fungus Candida albicans.J Antimicrob Chemother. 2020 Feb 1;75(2):257-270. doi: 10.1093/jac/dkz400. J Antimicrob Chemother. 2020. PMID: 31603213 Free PMC article. Review.
-
Genetic diversity and antifungal susceptibility profiles in causative agents of sporotrichosis.BMC Infect Dis. 2014 Apr 23;14:219. doi: 10.1186/1471-2334-14-219. BMC Infect Dis. 2014. PMID: 24755107 Free PMC article.
References
-
- Cahuzac B., Cerdan R., Felenbok B., Guittet E. 2001. The solution structure of an AlcR-DNA complex sheds light onto the unique tight and monomeric DNA binding of a Zn(2)Cys(6) protein. Structure 9:827–836 - PubMed
-
- Calabrese D., Bille J., Sanglard D. 2000. A novel multidrug efflux transporter gene of the major facilitator superfamily from Candida albicans (FLU1) conferring resistance to fluconazole. Microbiology 146:2743–2754 - PubMed
-
- Carroll J. S., et al. 2006. Genome-wide analysis of estrogen receptor binding sites. Nat. Genet. 38:1289–1297 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

