RAP80-directed tuning of BRCA1 homologous recombination function at ionizing radiation-induced nuclear foci
- PMID: 21406551
- PMCID: PMC3070932
- DOI: 10.1101/gad.2011011
RAP80-directed tuning of BRCA1 homologous recombination function at ionizing radiation-induced nuclear foci
Abstract
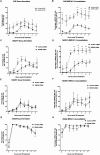
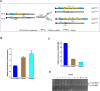
In response to DNA double-strand breaks (DSBs), BRCA1 forms biochemically distinct complexes with certain other DNA damage response proteins. These structures, some of which are required for homologous recombination (HR)-type DSB repair, concentrate at distinct nuclear foci that demarcate sites of genome breakage. Polyubiquitin binding by one of these structures, the RAP80/BRCA1 complex, is required for efficient BRCA1 focal recruitment, but the relationship of this process to the execution of HR has been unclear. We found that this complex actively suppresses otherwise exaggerated, BRCA1-driven HR. By controlling the kinetics by which other BRCA1-interacting proteins that promote HR concentrate together with BRCA1 in nuclear foci, RAP80/BRCA1 complexes suppress excessive DSB end processing, HR-type DSB repair, and overt chromosomal instability. Since chromosomal instability emerges when BRCA1 HR function is either unbridled or absent, active tuning of BRCA1 activity, executed in nuclear foci, is important to genome integrity maintenance.
Figures





References
-
- Branzei D, Vanoli F, Foiani M 2008. SUMOylation regulates Rad18-mediated template switch. Nature 456: 915–920 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
