Impulse control: temporal dynamics in gene transcription
- PMID: 21414481
- PMCID: PMC3148525
- DOI: 10.1016/j.cell.2011.02.015
Impulse control: temporal dynamics in gene transcription
Abstract
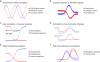
Regulatory circuits controlling gene expression constantly rewire to adapt to environmental stimuli, differentiation cues, and disease. We review our current understanding of the temporal dynamics of gene expression in eukaryotes and prokaryotes and the molecular mechanisms that shape them. We delineate several prototypical temporal patterns, including "impulse" (or single-pulse) patterns in response to transient environmental stimuli, sustained (or state-transitioning) patterns in response to developmental cues, and oscillating patterns. We focus on impulse responses and their higher-order temporal organization in regulons and cascades and describe how core protein circuits and cis-regulatory sequences in promoters integrate with chromatin architecture to generate these responses.
Copyright © 2011 Elsevier Inc. All rights reserved.
Figures

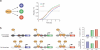

References
-
- Alon U. Network motifs: theory and experimental approaches. Nat Rev Genet. 2007;8:450–461. - PubMed
-
- Amit I, Citri A, Shay T, Lu Y, Katz M, Zhang F, Tarcic G, Siwak D, Lahad J, Jacob-Hirsch J, et al. A module of negative feedback regulators defines growth factor signaling. Nat Genet. 2007a;39:503–512. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

