MKS and NPHP modules cooperate to establish basal body/transition zone membrane associations and ciliary gate function during ciliogenesis
- PMID: 21422230
- PMCID: PMC3063147
- DOI: 10.1083/jcb.201012116
MKS and NPHP modules cooperate to establish basal body/transition zone membrane associations and ciliary gate function during ciliogenesis
Abstract
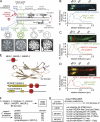
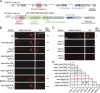
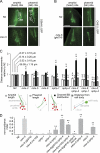
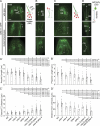
Meckel-Gruber syndrome (MKS), nephronophthisis (NPHP), and related ciliopathies present with overlapping phenotypes and display considerable allelism between at least twelve different genes of largely unexplained function. We demonstrate that the conserved C. elegans B9 domain (MKS-1, MKSR-1, and MKSR-2), MKS-3/TMEM67, MKS-5/RPGRIP1L, MKS-6/CC2D2A, NPHP-1, and NPHP-4 proteins exhibit essential, collective functions at the transition zone (TZ), an underappreciated region at the base of all cilia characterized by Y-shaped assemblages that link axoneme microtubules to surrounding membrane. These TZ proteins functionally interact as members of two distinct modules, which together contribute to an early ciliogenic event. Specifically, MKS/MKSR/NPHP proteins establish basal body/TZ membrane attachments before or coinciding with intraflagellar transport-dependent axoneme extension and subsequently restrict accumulation of nonciliary components within the ciliary compartment. Together, our findings uncover a unified role for eight TZ-localized proteins in basal body anchoring and establishing a ciliary gate during ciliogenesis, and suggest that disrupting ciliary gate function contributes to phenotypic features of the MKS/NPHP disease spectrum.
Figures



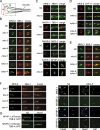


References
-
- Bialas N.J., Inglis P.N., Li C., Robinson J.F., Parker J.D., Healey M.P., Davis E.E., Inglis C.D., Toivonen T., Cottell D.C., et al. 2009. Functional interactions between the ciliopathy-associated Meckel syndrome 1 (MKS1) protein and two novel MKS1-related (MKSR) proteins. J. Cell Sci. 122:611–624 10.1242/jcs.028621 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

