Readers of histone modifications
- PMID: 21423274
- PMCID: PMC3131977
- DOI: 10.1038/cr.2011.42
Readers of histone modifications
Abstract
Histone modifications not only play important roles in regulating chromatin structure and nuclear processes but also can be passed to daughter cells as epigenetic marks. Accumulating evidence suggests that the key function of histone modifications is to signal for recruitment or activity of downstream effectors. Here, we discuss the latest discovery of histone-modification readers and how the modification language is interpreted.
Figures
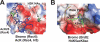
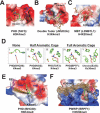


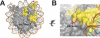

References
-
- Li B, Carey M, Workman JL. The role of chromatin during transcription. Cell. 2007;128:707–719. - PubMed
-
- Ahmad K, Henikoff S. Epigenetic consequences of nucleosome dynamics. Cell. 2002;111:281–284. - PubMed
-
- Kouzarides T. Chromatin modifications and their function. Cell. 2007;128:693–705. - PubMed
-
- Workman JL, Kingston RE. Alteration of nucleosome structure as a mechanism of transcriptional regulation. Annu Rev Biochem. 1998;67:545–579. - PubMed
-
- Shogren-Knaak M, Ishii H, Sun JM, Pazin MJ, Davie JR, Peterson CL. Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science. 2006;311:844–847. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

