Clues to evolution of the SERA multigene family in 18 Plasmodium species
- PMID: 21423628
- PMCID: PMC3058004
- DOI: 10.1371/journal.pone.0017775
Clues to evolution of the SERA multigene family in 18 Plasmodium species
Abstract
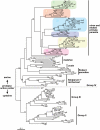
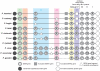
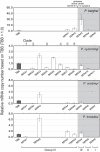
SERA gene sequences were newly determined from 11 primate Plasmodium species including two human parasites, P. ovale and P. malariae, and the evolutionary history of SERA genes was analyzed together with 7 known species. All have one each of Group I to III cysteine-type SERA genes and varying number of Group IV serine-type SERA genes in tandem cluster. Notably, Group IV SERA genes were ascertained in all mammalian parasite lineages; and in two primate parasite lineages gene events such as duplication, truncation, fragmentation and gene loss occurred at high frequency in a manner that mimics the birth-and-death evolution model. Transcription profile of individual SERA genes varied greatly among rodent and monkey parasites. Results support the lineage-specific evolution of the Plasmodium SERA gene family. These findings provide further impetus for studies that could clarify/provide proof-of-concept that duplications of SERA genes were associated with the parasites' expansion of host range and the evolutionary conundrums of multigene families in Plasmodium.
Conflict of interest statement
Figures



References
-
- Horii T, Shirai H, Jie L, Ishii KJ, Palacpac NQ, et al. Evidences of protection against blood-stage infection of Plasmodium falciparum by the novel protein vaccine SE36. Parasitol Int. 2010;593:380–386. - PubMed
-
- Okech BA, Nalunkuma A, Okello D, Pang XL, Suzue K, et al. Natural human immunoglobulin G subclass responses to Plasmodium falciparum serine repeat antigen in Uganda. Am J Trop Med Hyg. 2001;65:912–917. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

