Drosophila genes that affect meiosis duration are among the meiosis related genes that are more often found duplicated
- PMID: 21423746
- PMCID: PMC3053365
- DOI: 10.1371/journal.pone.0017512
Drosophila genes that affect meiosis duration are among the meiosis related genes that are more often found duplicated
Abstract
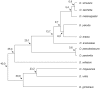
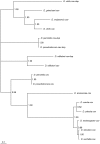
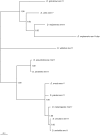
Using a phylogenetic approach, the examination of 33 meiosis/meiosis-related genes in 12 Drosophila species, revealed nine independent gene duplications, involving the genes cav, mre11, meiS332, polo and mtrm. Evidence is provided that at least eight out of the nine gene duplicates are functional. Therefore, the rate at which Drosophila meiosis/meiosis-related genes are duplicated and retained is estimated to be 0.0012 per gene per million years, a value that is similar to the average for all Drosophila genes. It should be noted that by using a phylogenetic approach the confounding effect of concerted evolution, that is known to lead to overestimation of the duplication and retention rate, is avoided. This is an important issue, since even in our moderate size sample, evidence for long-term concerted evolution (lasting for more than 30 million years) was found for the meiS332 gene pair in species of the Drosophila subgenus. Most striking, in contrast to theoretical expectations, is the finding that genes that encode proteins that must follow a close stoichiometric balance, such as polo, mtrm and meiS332 have been found duplicated. The duplicated genes may be examples of gene neofunctionalization. It is speculated that meiosis duration may be a trait that is under selection in Drosophila and that it has different optimal values in different species.
Conflict of interest statement
Figures
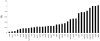

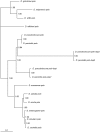
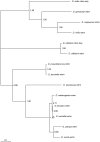
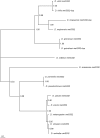
Similar articles
-
Neofunctionalization of young duplicate genes in Drosophila.Proc Natl Acad Sci U S A. 2013 Oct 22;110(43):17409-14. doi: 10.1073/pnas.1313759110. Epub 2013 Oct 7. Proc Natl Acad Sci U S A. 2013. PMID: 24101476 Free PMC article.
-
Testis- and ovary-expressed polo-like kinase transcripts and gene duplications affect male fertility when expressed in the Drosophila melanogaster germline.G3 (Bethesda). 2025 Jan 8;15(1):jkae273. doi: 10.1093/g3journal/jkae273. G3 (Bethesda). 2025. PMID: 39566185 Free PMC article.
-
Functional requirements driving the gene duplication in 12 Drosophila species.BMC Genomics. 2013 Aug 15;14:555. doi: 10.1186/1471-2164-14-555. BMC Genomics. 2013. PMID: 23945147 Free PMC article.
-
Theories for analyzing polymorphism data in duplicated genes.Genes Genet Syst. 2004 Apr;79(2):65-75. doi: 10.1266/ggs.79.65. Genes Genet Syst. 2004. PMID: 15215672 Review.
-
Evolution of the duplicated intracellular lipid-binding protein genes of teleost fishes.Mol Genet Genomics. 2017 Aug;292(4):699-727. doi: 10.1007/s00438-017-1313-5. Epub 2017 Apr 7. Mol Genet Genomics. 2017. PMID: 28389698 Review.
Cited by
-
Narya, a RING finger domain-containing protein, is required for meiotic DNA double-strand break formation and crossover maturation in Drosophila melanogaster.PLoS Genet. 2019 Jan 7;15(1):e1007886. doi: 10.1371/journal.pgen.1007886. eCollection 2019 Jan. PLoS Genet. 2019. PMID: 30615609 Free PMC article.
-
Functional Consequences of the Evolution of Matrimony, a Meiosis-Specific Inhibitor of Polo Kinase.Mol Biol Evol. 2019 Jan 1;36(1):69-83. doi: 10.1093/molbev/msy197. Mol Biol Evol. 2019. PMID: 30351378 Free PMC article.
-
Recurrent Innovation at Genes Required for Telomere Integrity in Drosophila.Mol Biol Evol. 2017 Feb 1;34(2):467-482. doi: 10.1093/molbev/msw248. Mol Biol Evol. 2017. PMID: 27836984 Free PMC article.
-
Evaluation of Beauveria bassiana infection in the hemolymph serum proteins of the housefly, Musca domestica L. (Diptera: Muscidae).Environ Sci Pollut Res Int. 2017 Nov;24(31):24714-24724. doi: 10.1007/s11356-017-0193-x. Epub 2017 Sep 21. Environ Sci Pollut Res Int. 2017. PMID: 28936573
-
Well-Annotated microRNAomes Do Not Evidence Pervasive miRNA Loss.Genome Biol Evol. 2018 Jun 1;10(6):1457-1470. doi: 10.1093/gbe/evy096. Genome Biol Evol. 2018. PMID: 29788279 Free PMC article.
References
-
- Lynch M, Conery JS. The evolutionary fate and consequences of duplicate genes. Science. 2000;290:1151–1155. - PubMed
-
- Gao LZ, Innan H. Very low gene duplication rate in the yeast genome. Science. 2004;306:1367–1370. - PubMed
-
- Papp B, Pal C, Hurst LD. Dosage sensitivity and the evolution of gene families in yeast. Nature. 2003;424:194–197. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

