Nuclear phospholipid scramblase 1 prolongs the mitotic expansion of granulocyte precursors during G-CSF-induced granulopoiesis
- PMID: 21447647
- PMCID: PMC3133439
- DOI: 10.1189/jlb.0111006
Nuclear phospholipid scramblase 1 prolongs the mitotic expansion of granulocyte precursors during G-CSF-induced granulopoiesis
Abstract
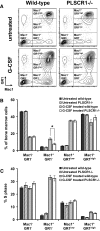
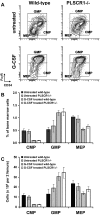
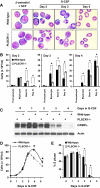
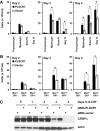
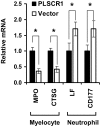
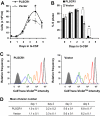
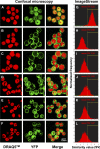
PLSCR1-/- mice exhibit normal, steady-state hematologic parameters but impaired emergency granulopoiesis upon in vivo administration of G-CSF. The mechanism by which PLSCR1 contributes to G-CSF-induced neutrophil production is largely unknown. We now report that the expansion of bone marrow myelocytes upon in vivo G-CSF treatment is reduced in PLSCR1-/- mice relative to WT. Using SCF-ER-Hoxb8-immortalized myeloid progenitors to examine the progression of G-CSF-driven granulocytic differentiation in vitro, we found that PLSCR1 prolongs the period of mitotic expansion of proliferative granulocyte precursors, thereby giving rise to increased neutrophil production from their progenitors. This effect of PLSCR1 is blocked by a ΔNLS-PLSCR1, which prevents its nuclear import. By contrast, mutation that prevents the membrane association of PLSCR1 has minimal impact on the role of PLSCR1 in G-CSF-induced granulopoiesis. These data imply that the capacity of PLSCR1 to augment G-CSF-dependent production of mature neutrophils from myeloid progenitors is unrelated to its reported activities at the endofacial surface of the plasma membrane but does require entry of the protein into the nucleus, suggesting that this response is mediated through the observed effects of PLSCR1 on gene transcription.
Figures

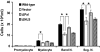
Comment in
-
Editorial: Scrambling for a response to G-CSF.J Leukoc Biol. 2011 Aug;90(2):215-6. doi: 10.1189/jlb.0311138. J Leukoc Biol. 2011. PMID: 21807746 No abstract available.
Similar articles
-
Normal hemostasis but defective hematopoietic response to growth factors in mice deficient in phospholipid scramblase 1.Blood. 2002 Jun 1;99(11):4030-8. doi: 10.1182/blood-2001-12-0271. Blood. 2002. PMID: 12010804
-
Evaluation of role of G-CSF in the production, survival, and release of neutrophils from bone marrow into circulation.Blood. 2002 Aug 1;100(3):854-61. doi: 10.1182/blood.v100.3.854. Blood. 2002. PMID: 12130495
-
Editorial: Scrambling for a response to G-CSF.J Leukoc Biol. 2011 Aug;90(2):215-6. doi: 10.1189/jlb.0311138. J Leukoc Biol. 2011. PMID: 21807746 No abstract available.
-
Phospholipid scramblase 1: a protein with multiple functions via multiple molecular interactors.Cell Commun Signal. 2022 Jun 1;20(1):78. doi: 10.1186/s12964-022-00895-3. Cell Commun Signal. 2022. PMID: 35650588 Free PMC article. Review.
-
Neutrophil Homeostasis and Emergency Granulopoiesis: The Example of Systemic Juvenile Idiopathic Arthritis.Front Immunol. 2021 Dec 13;12:766620. doi: 10.3389/fimmu.2021.766620. eCollection 2021. Front Immunol. 2021. PMID: 34966386 Free PMC article. Review.
Cited by
-
Beyond apoptosis: the mechanism and function of phosphatidylserine asymmetry in the membrane of activating mast cells.Bioarchitecture. 2014;4(4-5):127-37. doi: 10.1080/19490992.2014.995516. Bioarchitecture. 2014. PMID: 25759911 Free PMC article. Review.
-
Biophysical characterization of the DNA binding motif of human phospholipid scramblase 1.Eur Biophys J. 2022 Dec;51(7-8):579-593. doi: 10.1007/s00249-022-01621-0. Epub 2022 Oct 19. Eur Biophys J. 2022. PMID: 36260146
-
Phospholipid scramblase 1 amplifies anaphylactic reactions in vivo.PLoS One. 2017 Mar 10;12(3):e0173815. doi: 10.1371/journal.pone.0173815. eCollection 2017. PLoS One. 2017. PMID: 28282470 Free PMC article.
-
Phospholipid Scramblases: Role in Cancer Progression and Anticancer Therapeutics.Front Genet. 2022 Mar 29;13:875894. doi: 10.3389/fgene.2022.875894. eCollection 2022. Front Genet. 2022. PMID: 35422844 Free PMC article. Review.
-
Phospholipid scramblase 1 mediates type i interferon-induced protection against staphylococcal α-toxin.Cell Host Microbe. 2012 Jan 19;11(1):70-80. doi: 10.1016/j.chom.2011.12.004. Cell Host Microbe. 2012. PMID: 22264514 Free PMC article.
References
-
- Nicola N. A., Metcalf D., Matsumoto M., Johnson G. R. (1983) Purification of a factor inducing differentiation in murine myelomonocytic leukemia cells. Identification as granulocyte colony-stimulating factor. J. Biol. Chem. 258, 9017–9023 - PubMed
-
- Souza L. M., Boone T. C., Gabrilove J., Lai P. H., Zsebo K. M., Murdock D. C., Chazin V. R., Bruszewski J., Lu H., Chen K. K., et al. (1986) Recombinant human granulocyte colony-stimulating factor: effects on normal and leukemic myeloid cells. Science 232, 61–65 - PubMed
-
- Lieschke G. J., Grail D., Hodgson G., Metcalf D., Stanley E., Cheers C., Fowler K. J., Basu S., Zhan Y. F., Dunn A. R. (1994) Mice lacking granulocyte colony-stimulating factor have chronic neutropenia, granulocyte and macrophage progenitor cell deficiency, and impaired neutrophil mobilization. Blood 84, 1737–1746 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

