Dissociation by Pelota, Hbs1 and ABCE1 of mammalian vacant 80S ribosomes and stalled elongation complexes
- PMID: 21448132
- PMCID: PMC3101999
- DOI: 10.1038/emboj.2011.93
Dissociation by Pelota, Hbs1 and ABCE1 of mammalian vacant 80S ribosomes and stalled elongation complexes
Abstract
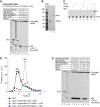
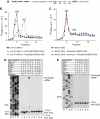
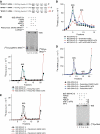
No-go decay (NGD) and non-stop decay (NSD) are eukaryotic surveillance mechanisms that target mRNAs on which elongation complexes (ECs) are stalled by, for example, stable secondary structures (NGD) or due to the absence of a stop codon (NSD). Two interacting proteins Dom34(yeast)/Pelota(mammals) and Hbs1, which are paralogues of eRF1 and eRF3, are implicated in these processes. Dom34/Hbs1 were shown to promote dissociation of stalled ECs and release of intact peptidyl-tRNA. Using an in vitro reconstitution approach, we investigated the activities of mammalian Pelota/Hbs1 and report that Pelota/Hbs1 also induced dissociation of ECs and release of peptidyl-tRNA, but only in the presence of ABCE1. Whereas Pelota and ABCE1 were essential, Hbs1 had a stimulatory effect. Importantly, ABCE1/Pelota/Hbs1 dissociated ECs containing only a limited number of mRNA nucleotides downstream of the P-site, which suggests that ABCE1/Pelota/Hbs1 would disassemble NSD complexes stalled at the 3'-end, but not pre-cleavage NGD complexes stalled in the middle of mRNA. ABCE1/Pelota/Hbs1 also dissociated vacant 80S ribosomes, which stimulated 48S complex formation, suggesting that Pelota/Hbs1 have an additional role outside of NGD.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures




References
-
- Acker MG, Kolitz SE, Mitchell SF, Nanda JS, Lorsch JR (2007) Reconstitution of yeast translation initiation. Methods Enzymol 430: 111–145 - PubMed
-
- Alkalaeva EZ, Pisarev AV, Frolova LY, Kisselev LL, Pestova TV (2006) In vitro reconstitution of eukaryotic translation reveals cooperativity between release factors eRF1 and eRF3. Cell 125: 1125–1136 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

