Genome-wide characterization of chromatin binding and nucleosome spacing activity of the nucleosome remodelling ATPase ISWI
- PMID: 21448136
- PMCID: PMC3102003
- DOI: 10.1038/emboj.2011.98
Genome-wide characterization of chromatin binding and nucleosome spacing activity of the nucleosome remodelling ATPase ISWI
Abstract
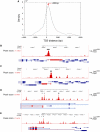
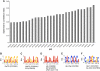
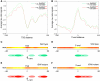
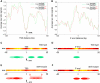
The evolutionarily conserved ATP-dependent nucleosome remodelling factor ISWI can space nucleosomes affecting a variety of nuclear processes. In Drosophila, loss of ISWI leads to global transcriptional defects and to dramatic alterations in higher-order chromatin structure, especially on the male X chromosome. In order to understand if chromatin condensation and gene expression defects, observed in ISWI mutants, are directly correlated with ISWI nucleosome spacing activity, we conducted a genome-wide survey of ISWI binding and nucleosome positioning in wild-type and ISWI mutant chromatin. Our analysis revealed that ISWI binds both genic and intergenic regions. Remarkably, we found that ISWI binds genes near their promoters causing specific alterations in nucleosome positioning at the level of the Transcription Start Site, providing an important insights in understanding ISWI role in higher eukaryote transcriptional regulation. Interestingly, differences in nucleosome spacing, between wild-type and ISWI mutant chromatin, tend to accumulate on the X chromosome for all ISWI-bound genes analysed. Our study shows how in higher eukaryotes the activity of the evolutionarily conserved nucleosome remodelling factor ISWI regulates gene expression and chromosome organization genome-wide.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures



References
-
- Burgio G, La Rocca G, Sala A, Arancio W, Di Gesu D, Collesano M, Sperling AS, Armstrong JA, van Heeringen SJ, Logie C, Tamkun JW, Corona DF (2008) Genetic identification of a network of factors that functionally interact with the nucleosome remodeling ATPase ISWI. PLoS Genet 4: e1000089. - PMC - PubMed
-
- Corona DF, Armstrong JA, Tamkun JW (2004) Genetic and cytological analysis of Drosophila chromatin-remodeling factors. Methods Enzymol 377: 70–85 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

