Point mutations in the Rpb9-homologous domain of Rpc11 that impair transcription termination by RNA polymerase III
- PMID: 21450810
- PMCID: PMC3152337
- DOI: 10.1093/nar/gkr182
Point mutations in the Rpb9-homologous domain of Rpc11 that impair transcription termination by RNA polymerase III
Abstract
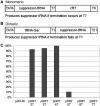
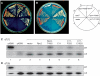
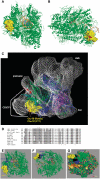
RNA polymerase III recognizes and pauses at its terminator, an oligo(dT) tract in non-template DNA, terminates 3' oligo(rU) synthesis within this sequence, and releases the RNA. The pol III subunit Rpc11p (C11) mediates RNA 3'-5' cleavage in the catalytic center of pol III during pausing. The amino and carboxyl regions of C11 are homologous to domains of the pol II subunit Rpb9p, and the pol II elongation and RNA cleavage factor, TFIIS, respectively. We isolated C11 mutants from Schizosaccharomyces pombe that cause pol III to readthrough terminators in vivo. Mutant RNA confirmed the presence of terminator readthrough transcripts. A predominant mutation site, F32, resides in the C11 Rpb9-like domain. Another mutagenic approach confirmed the F32 mutation and also isolated I34 and Y30 mutants. Modeling Y30, F32 and I34 of C11 in available cryoEM pol III structures predicts a hydrophobic patch that may interface with C53/37. Another termination mutant, Rpc2-T455I, appears to reside internally, near the RNA-DNA hybrid. We show that the Rpb9 and TFIIS homologous mutants of C11 reflect distinct activities, that differentially affect terminator recognition and RNA 3' cleavage. We propose that these C11 domains integrate action at the upper jaw and center of pol III during termination.
Figures



Similar articles
-
Transcription termination by the eukaryotic RNA polymerase III.Biochim Biophys Acta. 2013 Mar-Apr;1829(3-4):318-30. doi: 10.1016/j.bbagrm.2012.10.006. Epub 2012 Oct 23. Biochim Biophys Acta. 2013. PMID: 23099421 Free PMC article. Review.
-
RNA polymerase III mutants in TFIIFα-like C37 that cause terminator readthrough with no decrease in transcription output.Nucleic Acids Res. 2013 Jan 7;41(1):139-55. doi: 10.1093/nar/gks985. Epub 2012 Oct 23. Nucleic Acids Res. 2013. PMID: 23093604 Free PMC article.
-
The RNA cleavage activity of RNA polymerase III is mediated by an essential TFIIS-like subunit and is important for transcription termination.Genes Dev. 1998 Dec 15;12(24):3857-71. doi: 10.1101/gad.12.24.3857. Genes Dev. 1998. PMID: 9869639 Free PMC article.
-
Active Center Control of Termination by RNA Polymerase III and tRNA Gene Transcription Levels In Vivo.PLoS Genet. 2016 Aug 12;12(8):e1006253. doi: 10.1371/journal.pgen.1006253. eCollection 2016 Aug. PLoS Genet. 2016. PMID: 27518095 Free PMC article.
-
Comparison of the RNA polymerase III transcription machinery in Schizosaccharomyces pombe, Saccharomyces cerevisiae and human.Nucleic Acids Res. 2001 Jul 1;29(13):2675-90. doi: 10.1093/nar/29.13.2675. Nucleic Acids Res. 2001. PMID: 11433012 Free PMC article. Review.
Cited by
-
Transcription termination by the eukaryotic RNA polymerase III.Biochim Biophys Acta. 2013 Mar-Apr;1829(3-4):318-30. doi: 10.1016/j.bbagrm.2012.10.006. Epub 2012 Oct 23. Biochim Biophys Acta. 2013. PMID: 23099421 Free PMC article. Review.
-
Specialization versus conservation: How Pol I and Pol III use the conserved architecture of the pre-initiation complex for specialized transcription.Transcription. 2016 Aug 7;7(4):127-32. doi: 10.1080/21541264.2016.1203628. Epub 2016 Jun 21. Transcription. 2016. PMID: 27327079 Free PMC article. Review.
-
Structural insights into nuclear transcription by eukaryotic DNA-dependent RNA polymerases.Nat Rev Mol Cell Biol. 2022 Sep;23(9):603-622. doi: 10.1038/s41580-022-00476-9. Epub 2022 May 3. Nat Rev Mol Cell Biol. 2022. PMID: 35505252 Review.
-
A methods review on use of nonsense suppression to study 3' end formation and other aspects of tRNA biogenesis.Gene. 2015 Feb 1;556(1):35-50. doi: 10.1016/j.gene.2014.11.034. Epub 2014 Nov 18. Gene. 2015. PMID: 25447915 Free PMC article. Review.
-
RNA polymerase III subunits C37/53 modulate rU:dA hybrid 3' end dynamics during transcription termination.Nucleic Acids Res. 2019 Jan 10;47(1):310-327. doi: 10.1093/nar/gky1109. Nucleic Acids Res. 2019. PMID: 30407541 Free PMC article.
References
-
- Geiger SR, Lorenzen K, Schreieck A, Hanecker P, Kostrewa D, Heck AJ, Cramer P. RNA polymerase I contains a TFIIF-related DNA-binding subcomplex. Mol. Cell. 2010;39:583–594. - PubMed
-
- Vannini A, Ringel R, Kusser AG, Berninghausen O, Kassavetis GA, Cramer P. Molecular basis of RNA polymerase III transcription repression by Maf1. Cell. 2010;143:59–70. - PubMed
-
- Dieci G, Sentenac A. Facilitated recycling by RNA polymerase III. Cell. 1996;84:245–252. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

