Shotgun proteomics aids discovery of novel protein-coding genes, alternative splicing, and "resurrected" pseudogenes in the mouse genome
- PMID: 21460061
- PMCID: PMC3083093
- DOI: 10.1101/gr.114272.110
Shotgun proteomics aids discovery of novel protein-coding genes, alternative splicing, and "resurrected" pseudogenes in the mouse genome
Abstract
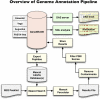
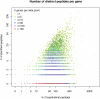
Recent advances in proteomic mass spectrometry (MS) offer the chance to marry high-throughput peptide sequencing to transcript models, allowing the validation, refinement, and identification of new protein-coding loci. We present a novel pipeline that integrates highly sensitive and statistically robust peptide spectrum matching with genome-wide protein-coding predictions to perform large-scale gene validation and discovery in the mouse genome for the first time. In searching an excess of 10 million spectra, we have been able to validate 32%, 17%, and 7% of all protein-coding genes, exons, and splice boundaries, respectively. Moreover, we present strong evidence for the identification of multiple alternatively spliced translations from 53 genes and have uncovered 10 entirely novel protein-coding genes, which are not covered in any mouse annotation data sources. One such novel protein-coding gene is a fusion protein that spans the Ins2 and Igf2 loci to produce a transcript encoding the insulin II and the insulin-like growth factor 2-derived peptides. We also report nine processed pseudogenes that have unique peptide hits, demonstrating, for the first time, that they are not just transcribed but are translated and are therefore resurrected into new coding loci. This work not only highlights an important utility for MS data in genome annotation but also provides unique insights into the gene structure and propagation in the mouse genome. All these data have been subsequently used to improve the publicly available mouse annotation available in both the Vega and Ensembl genome browsers (http://vega.sanger.ac.uk).
Figures







References
-
- Abbott A 2010. Mouse project to find each gene's role: International Mouse Phenotyping Consortium launches with a massive funding commitment. Nature 465: 410 doi: 10.1038/465410a - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous
