The human gut mobile metagenome: a metazoan perspective
- PMID: 21468227
- PMCID: PMC3056110
- DOI: 10.4161/gmic.1.6.14087
The human gut mobile metagenome: a metazoan perspective
Abstract
Using the culture independent TRACA system in conjunction with a comparative metagenomic approach, we have recently explored the pool of plasmids associated with the human gut mobile metagenome. This revealed that some plasmids or plasmid families are present in the gut microbiomes of geographically isolated human hosts with a broad global distribution (America, Japan and Europe), and are potentially unique to the human gut microbiome. Functions encoded by the most widely distributed plasmid (pTRACA22) were found to be enriched in the human gut microbiome when compared to microbial communities from other environments, and of particular interest was the increased prevalence of a putative RelBE toxin-antitoxin (TA) addiction module. Subsequent analysis revealed that this was most closely related to putative TA modules from gut associated bacteria belonging to the Firmicutes, but homologues of the RelE toxin were associated with all major bacterial divisions comprising the human gut microbiota. In this addendum, functions of the gut mobile metagenome are considered from the perspective of the human host, and within the context of the hologenome theory of human evolution. In doing so, our original analysis is also extended to include the gut metagenomes of a further 124 individuals comprising the METAHIT dataset. Differences in the incidence and relative abundance of pTRACA22 and associated TA modules between healthy individuals and those with inflammatory bowel diseases are explored, and potential functions of pTRACA22 type RelBE modules in the human gut microbiome are discussed.
Keywords: gut microbiota; holobiont; hologenome; horizontal gene transfer; metagenomics; mobile genetic elements; mobile metagenome; toxin-antitoxin addiction module.
© 2010 Landes Bioscience
Figures
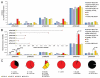

Comment on
-
Comparative metagenomic analysis of plasmid encoded functions in the human gut microbiome.BMC Genomics. 2010 Jan 19;11:46. doi: 10.1186/1471-2164-11-46. BMC Genomics. 2010. PMID: 20085629 Free PMC article.
References
-
- Bäckhed F, Ley RE, Sonnenburg JL, Peterson DA, Gordon JI. Host-bacterial mutualism in the human intestine. Science. 2005;307:1915–1920. - PubMed
-
- Zilber-Rosenberg I, Rosenberg E. Role of microorganisms in the evolution of animals and plants: the hologenome theory of evolution. FEMS Microbiol Rev. 2008;32:723–725. - PubMed
-
- Rosenberg E, Sharon G, Zilber-Rosenberg I. The hologenome theory of evolution contains Lamarckian aspects within a Darwinian framework. Environ Microbiol. 2009;11:2959–2962. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
