Solution structure of the monovalent lectin microvirin in complex with Man(alpha)(1-2)Man provides a basis for anti-HIV activity with low toxicity
- PMID: 21471192
- PMCID: PMC3121468
- DOI: 10.1074/jbc.M111.232678
Solution structure of the monovalent lectin microvirin in complex with Man(alpha)(1-2)Man provides a basis for anti-HIV activity with low toxicity
Abstract
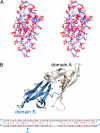
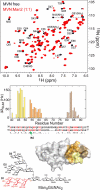
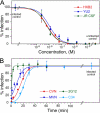
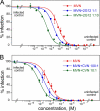
Lectins that bind surface envelope glycoprotein gp120 of HIV with high avidity can potently inhibit viral entry. Yet properties such as multivalency that facilitate strong interactions can also cause nonspecific binding and toxicity. The cyanobacterial lectin microvirin (MVN) is unusual as it potently inhibits HIV-1 with negligible toxicity compared with cyanovirin-N (CVN), its well studied antiviral homolog. To understand the structural and mechanistic basis for these differences, we solved the solution structure of MVN free and in complex with its ligand Manα(1-2)Man, and we compared specificity and time windows of inhibition with CVN and Manα(1-2)Man-specific mAb 2G12. We show by NMR and analytical ultracentrifugation that MVN is monomeric in solution, and we demonstrate by NMR that Manα(1-2)Man-terminating carbohydrates interact with a single carbohydrate-binding site. Synchronized infectivity assays show that 2G12, MVN, and CVN inhibit entry with distinct kinetics. Despite shared specificity for Manα(1-2)Man termini, combinations of the inhibitors are synergistic suggesting they recognize discrete glycans and/or dynamic glycan conformations on gp120. Entry assays employing amphotropic viruses show that MVN is inactive, whereas CVN potently inhibits both. In addition to demonstrating that HIV-1 can be inhibited through monovalent interactions, given the similarity of the carbohydrate-binding site common to MVN and CVN, these data suggest that gp120 behaves as a clustered glycan epitope and that multivalent-protein interactions achievable with CVN but not MVN are required for inhibition of some viruses.
Figures


References
-
- Taiwo B., Hicks C., Eron J. (2010) J. Antimicrob. Chemother. 65, 1100–1107 - PubMed
-
- Folkers G. K., Fauci A. S. (2010) JAMA 304, 350–351 - PubMed
-
- Chan D. C., Kim P. S. (1998) Cell 93, 681–684 - PubMed
-
- Leonard C. K., Spellman M. W., Riddle L., Harris R. J., Thomas J. N., Gregory T. J. (1990) J. Biol. Chem. 265, 10373–10382 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

