Differential genome-wide profiling of tandem 3' UTRs among human breast cancer and normal cells by high-throughput sequencing
- PMID: 21474764
- PMCID: PMC3083091
- DOI: 10.1101/gr.115295.110
Differential genome-wide profiling of tandem 3' UTRs among human breast cancer and normal cells by high-throughput sequencing
Abstract
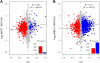
Tandem 3' UTRs produced by alternative polyadenylation (APA) play an important role in gene expression by impacting mRNA stability, translation, and translocation in cells. Several studies have investigated APA site switching in various physiological states; nevertheless, they only focused on either the genes with two known APA sites or several candidate genes. Here, we developed a strategy to study APA sites in a genome-wide fashion with second-generation sequencing technology which could not only identify new polyadenylation sites but also analyze the APA site switching of all genes, especially those with more than two APA sites. We used this strategy to explore the profiling of APA sites in two human breast cancer cell lines, MCF7 and MB231, and one cultured mammary epithelial cell line, MCF10A. More than half of the identified polyadenylation sites are not included in human poly(A) databases. While MCF7 showed shortening 3' UTRs, more genes in MB231 switched to distal poly(A) sites. Several gene ontology (GO) terms and pathways were enriched in the list of genes with switched APA sites, including cell cycle, apoptosis, and metabolism. These results suggest a more complex regulation of APA sites in cancer cells than previously thought. In short, our novel unbiased method can be a powerful approach to cost-effectively investigate the complex mechanism of 3' UTR switching in a genome-wide fashion among various physiological processes and diseases.
Figures


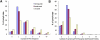
References
-
- Agresti A 2002. Categorical data analysis. Wiley-Interscience, New York
-
- Aptsiauri N, Cabrera T, Garcia-Lora A, Lopez-Nevot MA, Ruiz-Cabello F, Garrido F 2007a. MHC class I antigens and immune surveillance in transformed cells. Int Rev Cytol 256: 139–189 - PubMed
-
- Aptsiauri N, Cabrera T, Mendez R, Garcia-Lora A, Ruiz-Cabello F, Garrido F 2007b. Role of altered expression of HLA class I molecules in cancer progression. Adv Exp Med Biol 601: 123–131 - PubMed
-
- Bertone P, Stolc V, Royce TE, Rozowsky JS, Urban AE, Zhu X, Rinn JL, Tongprasit W, Samanta M, Weissman S, et al. 2004. Global identification of human transcribed sequences with genome tiling arrays. Science 306: 2242–2246 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
