A proteomic and cellular analysis of uropods in the pathogen Entamoeba histolytica
- PMID: 21483708
- PMCID: PMC3071361
- DOI: 10.1371/journal.pntd.0001002
A proteomic and cellular analysis of uropods in the pathogen Entamoeba histolytica
Abstract
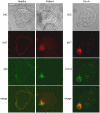
Exposure of Entamoeba histolytica to specific ligands induces cell polarization via the activation of signalling pathways and cytoskeletal elements. The process leads to formation of a protruding pseudopod at the front of the cell and a retracting uropod at the rear. In the present study, we show that the uropod forms during the exposure of trophozoites to serum isolated from humans suffering of amoebiasis. To investigate uropod assembly, we used LC-MS/MS technology to identify protein components in isolated uropod fractions. The galactose/N-acetylgalactosamine lectin, the immunodominant antigen M17 (which is specifically recognized by serum from amoeba-infected persons) and a few other cells adhesion-related molecules were primarily involved. Actin-rich cytoskeleton components, GTPases from the Rac and Rab families, filamin, α-actinin and a newly identified ezrin-moesin-radixin protein were the main factors found to potentially interact with capped receptors. A set of specific cysteine proteases and a serine protease were enriched in isolated uropod fractions. However, biological assays indicated that cysteine proteases are not involved in uropod formation in E. histolytica, a fact in contrast to the situation in human motile immune cells. The surface proteins identified here are testable biomarkers which may be either recognized by the immune system and/or released into the circulation during amoebiasis.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

Similar articles
-
Proteomic profiling reveals that EhPC4 transcription factor induces cell migration through up-regulation of the 16-kDa actin-binding protein EhABP16 in Entamoeba histolytica.J Proteomics. 2014 Dec 5;111:46-58. doi: 10.1016/j.jprot.2014.03.041. Epub 2014 Apr 8. J Proteomics. 2014. PMID: 24721673
-
Chemotaxis of Entamoeba histolytica towards the pro-inflammatory cytokine TNF is based on PI3K signalling, cytoskeleton reorganization and the Galactose/N-acetylgalactosamine lectin activity.Cell Microbiol. 2008 Aug;10(8):1676-86. doi: 10.1111/j.1462-5822.2008.01158.x. Epub 2008 Apr 16. Cell Microbiol. 2008. PMID: 18419774
-
Cell polarization and adhesion in a motile pathogenic protozoan: role and fate of the Entamoeba histolytica Gal/GalNAc lectin.Microbes Infect. 2000 May;2(6):643-9. doi: 10.1016/s1286-4579(00)00361-0. Microbes Infect. 2000. PMID: 10884615 Review.
-
Signalization and cytoskeleton activity through myosin IB during the early steps of phagocytosis in Entamoeba histolytica: a proteomic approach.Cell Microbiol. 2005 Oct;7(10):1504-18. doi: 10.1111/j.1462-5822.2005.00573.x. Cell Microbiol. 2005. PMID: 16153248
-
New insights into molecular mechanisms of phagocytosis in Entamoeba histolytica by proteomic analysis.Arch Med Res. 2006 Feb;37(2):244-52. doi: 10.1016/j.arcmed.2005.10.003. Arch Med Res. 2006. PMID: 16380325 Review.
Cited by
-
Proteomic analysis of the cyst stage of Entamoeba histolytica.PLoS Negl Trop Dis. 2012;6(5):e1643. doi: 10.1371/journal.pntd.0001643. Epub 2012 May 8. PLoS Negl Trop Dis. 2012. PMID: 22590659 Free PMC article.
-
Fibronectin induces a transition from amoeboid to a fan morphology and modifies migration in Entamoeba histolytica.PLoS Pathog. 2024 Jul 25;20(7):e1012392. doi: 10.1371/journal.ppat.1012392. eCollection 2024 Jul. PLoS Pathog. 2024. PMID: 39052670 Free PMC article.
-
Overexpression of specific cysteine peptidases confers pathogenicity to a nonpathogenic Entamoeba histolytica clone.mBio. 2013 Mar 26;4(2):e00072-13. doi: 10.1128/mBio.00072-13. mBio. 2013. PMID: 23532975 Free PMC article.
-
A novel extracellular metallopeptidase domain shared by animal host-associated mutualistic and pathogenic microbes.PLoS One. 2012;7(1):e30287. doi: 10.1371/journal.pone.0030287. Epub 2012 Jan 27. PLoS One. 2012. PMID: 22299034 Free PMC article.
-
Localization of phosphatidylinositol 4,5-bisphosphate to lipid rafts and uroids in the human protozoan parasite Entamoeba histolytica.Infect Immun. 2013 Jun;81(6):2145-55. doi: 10.1128/IAI.00040-13. Epub 2013 Apr 1. Infect Immun. 2013. PMID: 23545298 Free PMC article.
References
-
- Bretscher MS. Getting membrane flow and the cytoskeleton to cooperate in moving cells. Cell. 1996;87:601–606. - PubMed
-
- Bretscher MS. Recap on cell migration. Traffic. 2008;9:198–199. - PubMed
-
- Sanchez-Madrid F, Serrador JM. Bringing up the rear: defining the roles of the uropod. Nat Rev Mol Cell Biol. 2009;10:353–359. - PubMed
-
- Haque R, Huston CD, Hughes M, Houpt E, Petri WA., Jr Amebiasis. N Engl J Med. 2003;348:1565–1573. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

