Metabolite cross-feeding enhances virulence in a model polymicrobial infection
- PMID: 21483753
- PMCID: PMC3069116
- DOI: 10.1371/journal.ppat.1002012
Metabolite cross-feeding enhances virulence in a model polymicrobial infection
Abstract
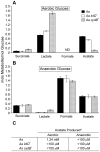
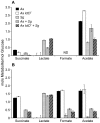
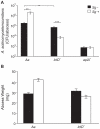
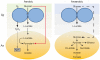
Microbes within polymicrobial infections often display synergistic interactions resulting in enhanced pathogenesis; however, the molecular mechanisms governing these interactions are not well understood. Development of model systems that allow detailed mechanistic studies of polymicrobial synergy is a critical step towards a comprehensive understanding of these infections in vivo. In this study, we used a model polymicrobial infection including the opportunistic pathogen Aggregatibacter actinomycetemcomitans and the commensal Streptococcus gordonii to examine the importance of metabolite cross-feeding for establishing co-culture infections. Our results reveal that co-culture with S. gordonii enhances the pathogenesis of A. actinomycetemcomitans in a murine abscess model of infection. Interestingly, the ability of A. actinomycetemcomitans to utilize L-lactate as an energy source is essential for these co-culture benefits. Surprisingly, inactivation of L-lactate catabolism had no impact on mono-culture growth in vitro and in vivo suggesting that A. actinomycetemcomitans L-lactate catabolism is only critical for establishing co-culture infections. These results demonstrate that metabolite cross-feeding is critical for A. actinomycetemcomitans to persist in a polymicrobial infection with S. gordonii supporting the idea that the metabolic properties of commensal bacteria alter the course of pathogenesis in polymicrobial communities.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


References
-
- Jenkinson HF, Lamont RJ. Oral microbial communities in sickness and in health. Trends Microbiol. 2005;13:589–595. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

