Genomic size of CENP-A domain is proportional to total alpha satellite array size at human centromeres and expands in cancer cells
- PMID: 21484447
- PMCID: PMC3565466
- DOI: 10.1007/s10577-011-9208-5
Genomic size of CENP-A domain is proportional to total alpha satellite array size at human centromeres and expands in cancer cells
Abstract
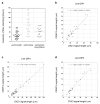
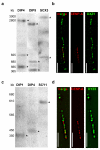
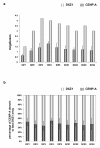
Human centromeres contain multi-megabase-sized arrays of alpha satellite DNA, a family of satellite DNA repeats based on a tandemly arranged 171 bp monomer. The centromere-specific histone protein CENP-A is assembled on alpha satellite DNA within the primary constriction, but does not extend along its entire length. CENP-A domains have been estimated to extend over 2,500 kb of alpha satellite DNA. However, these estimates do not take into account inter-individual variation in alpha satellite array sizes on homologous chromosomes and among different chromosomes. We defined the genomic distance of CENP-A chromatin on human chromosomes X and Y from different individuals. CENP-A chromatin occupied different genomic intervals on different chromosomes, but despite inter-chromosomal and inter-individual array size variation, the ratio of CENP-A to total alpha satellite DNA size remained consistent. Changes in the ratio of alpha satellite array size to CENP-A domain size were observed when CENP-A was overexpressed and when primary cells were transformed by disrupting interactions between the tumor suppressor protein Rb and chromatin. Our data support a model for centromeric domain organization in which the genomic limits of CENP-A chromatin varies on different human chromosomes, and imply that alpha satellite array size may be a more prominent predictor of CENP-A incorporation than chromosome size. In addition, our results also suggest that cancer transformation and amounts of centromeric heterochromatin have notable effects on the amount of alpha satellite that is associated with CENP-A chromatin.
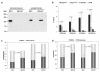
Figures
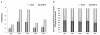
References
-
- ABRUZZO MA, GRIFFIN DK, MILLIE EA, SHEEAN LA, HASSOLD TJ. The effect of Y-chromosome alpha-satellite array length on the rate of sex chromosome disomy in human sperm. Hum Genet. 1996;97:819–23. - PubMed
-
- ALONSO A, MAHMOOD R, LI S, CHEUNG F, YODA K, WARBURTON PE. Genomic microarray analysis reveals distinct locations for the CENP-A binding domains in three human chromosome 13q32 neocentromeres. Hum Mol Genet. 2003;12:2711–21. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

