Chromatin globules: a common motif of higher order chromosome structure?
- PMID: 21489772
- PMCID: PMC3109114
- DOI: 10.1016/j.ceb.2011.03.009
Chromatin globules: a common motif of higher order chromosome structure?
Abstract
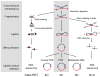
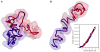
Recent technological advances in the field of chromosome conformation capture are facilitating tremendous progress in the ability to map the three-dimensional (3D) organization of chromosomes at a resolution of several Kb and at the scale of complete genomes. Here we review progress in analyzing chromosome organization in human cells by building 3D models of chromatin based on comprehensive chromatin interaction datasets. We describe recent experiments that suggest that long-range interactions between active functional elements are sufficient to drive folding of local chromatin domains into compact globular states. We propose that chromatin globules are commonly formed along chromosomes, in a cell type specific pattern, as a result of frequent long-range interactions among active genes and nearby regulatory elements. Further, we speculate that increasingly longer range interactions can drive aggregation of groups of globular domains. This process would yield a compartmentalized chromosome conformation, consistent with recent observations obtained with genome-wide chromatin interaction mapping.
Copyright © 2011 Elsevier Ltd. All rights reserved.
Figures

Similar articles
-
A (3D-Nuclear) Space Odyssey: Making Sense of Hi-C Maps.Genes (Basel). 2019 May 29;10(6):415. doi: 10.3390/genes10060415. Genes (Basel). 2019. PMID: 31146487 Free PMC article. Review.
-
Reconstruction of 3D genome architecture via a two-stage algorithm.BMC Bioinformatics. 2015 Nov 9;16:373. doi: 10.1186/s12859-015-0799-2. BMC Bioinformatics. 2015. PMID: 26553003 Free PMC article.
-
Graph rigidity reveals well-constrained regions of chromosome conformation embeddings.BMC Bioinformatics. 2012 Sep 21;13:241. doi: 10.1186/1471-2105-13-241. BMC Bioinformatics. 2012. PMID: 22998471 Free PMC article.
-
Mapping cis- and trans- chromatin interaction networks using chromosome conformation capture (3C).Methods Mol Biol. 2009;464:105-21. doi: 10.1007/978-1-60327-461-6_7. Methods Mol Biol. 2009. PMID: 18951182 Free PMC article.
-
Polycomb silencing: from linear chromatin domains to 3D chromosome folding.Curr Opin Genet Dev. 2014 Apr;25:30-7. doi: 10.1016/j.gde.2013.11.016. Epub 2014 Jan 14. Curr Opin Genet Dev. 2014. PMID: 24434548 Review.
Cited by
-
Analysis of long-range chromatin interactions using Chromosome Conformation Capture.Methods. 2012 Nov;58(3):192-203. doi: 10.1016/j.ymeth.2012.07.022. Epub 2012 Aug 15. Methods. 2012. PMID: 22903059 Free PMC article.
-
The Drosophila MI-2 chromatin-remodeling factor regulates higher-order chromatin structure and cohesin dynamics in vivo.PLoS Genet. 2012;8(8):e1002878. doi: 10.1371/journal.pgen.1002878. Epub 2012 Aug 9. PLoS Genet. 2012. PMID: 22912596 Free PMC article.
-
The Impact of Polycomb Group Proteins on 3D Chromatin Structure and Environmental Stresses in Plants.Plants (Basel). 2025 Mar 27;14(7):1038. doi: 10.3390/plants14071038. Plants (Basel). 2025. PMID: 40219106 Free PMC article. Review.
-
Probing long-range interactions by extracting free energies from genome-wide chromosome conformation capture data.BMC Bioinformatics. 2015 May 23;16:171. doi: 10.1186/s12859-015-0584-2. BMC Bioinformatics. 2015. PMID: 26001583 Free PMC article.
-
Exploring the three-dimensional organization of genomes: interpreting chromatin interaction data.Nat Rev Genet. 2013 Jun;14(6):390-403. doi: 10.1038/nrg3454. Epub 2013 May 9. Nat Rev Genet. 2013. PMID: 23657480 Free PMC article. Review.
References
-
- Dekker J, et al. Capturing Chromosome Conformation. Science. 2002;295(5558):1306–1311. - PubMed
-
- Simonis M, et al. Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C) Nat Genet. 2006;38(11):1348–1354. - PubMed
-
- Zhao Z, et al. Circular chromosome conformation capture (4C) uncovers extensive networks of epigenetically regulated intra- and interchromosomal interactions. Nat Genet. 2006;38(11):1341–1347. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

