Integrating the intrinsic conformational preferences of noncoded α-amino acids modified at the peptide bond into the noncoded amino acids database
- PMID: 21491493
- PMCID: PMC3092812
- DOI: 10.1002/prot.23009
Integrating the intrinsic conformational preferences of noncoded α-amino acids modified at the peptide bond into the noncoded amino acids database
Abstract
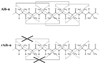
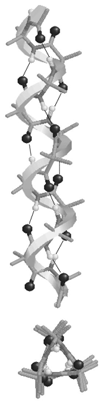
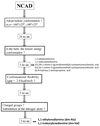
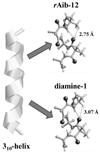
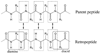
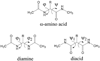
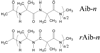
Recently, we reported a database (Noncoded Amino acids Database; http://recerca.upc.edu/imem/index.htm) that was built to compile information about the intrinsic conformational preferences of nonproteinogenic residues determined by quantum mechanical calculations, as well as bibliographic information about their synthesis, physical and spectroscopic characterization, the experimentally established conformational propensities, and applications (Revilla-López et al., J Phys Chem B 2010;114:7413-7422). The database initially contained the information available for α-tetrasubstituted α-amino acids. In this work, we extend NCAD to three families of compounds, which can be used to engineer peptides and proteins incorporating modifications at the--NHCO--peptide bond. Such families are: N-substituted α-amino acids, thio-α-amino acids, and diamines and diacids used to build retropeptides. The conformational preferences of these compounds have been analyzed and described based on the information captured in the database. In addition, we provide an example of the utility of the database and of the compounds it compiles in protein and peptide engineering. Specifically, the symmetry of a sequence engineered to stabilize the 3(10)-helix with respect to the α-helix has been broken without perturbing significantly the secondary structure through targeted replacements using the information contained in the database.
© 2011 Wiley-Liss, Inc.
Figures



Similar articles
-
NCAD, a database integrating the intrinsic conformational preferences of non-coded amino acids.J Phys Chem B. 2010 Jun 3;114(21):7413-22. doi: 10.1021/jp102092m. J Phys Chem B. 2010. PMID: 20455555 Free PMC article.
-
Comparison between the phi distribution of the amino acids in the protein database and NMR data indicates that amino acids have various phi propensities in the random coil conformation.J Mol Biol. 1995 Nov 24;254(2):322-33. doi: 10.1006/jmbi.1995.0619. J Mol Biol. 1995. PMID: 7490751
-
Noncoded Amino Acids in de Novo Metalloprotein Design: Controlling Coordination Number and Catalysis.Acc Chem Res. 2019 May 21;52(5):1160-1167. doi: 10.1021/acs.accounts.9b00032. Epub 2019 Apr 1. Acc Chem Res. 2019. PMID: 30933479 Free PMC article.
-
Noncoded amino acids in protein engineering: Structure-activity relationship studies of hirudin-thrombin interaction.Biotechnol Appl Biochem. 2018 Jan;65(1):69-80. doi: 10.1002/bab.1632. Epub 2018 Jan 16. Biotechnol Appl Biochem. 2018. PMID: 29230873 Review.
-
The twists and turns of beta-peptides.J Pept Res. 1999 Sep;54(3):206-17. doi: 10.1034/j.1399-3011.1999.00131.x. J Pept Res. 1999. PMID: 10517158 Review.
Cited by
-
Sequence-based prediction of the intrinsic solubility of peptides containing non-natural amino acids.Nat Commun. 2023 Nov 17;14(1):7475. doi: 10.1038/s41467-023-42940-w. Nat Commun. 2023. PMID: 37978172 Free PMC article.
-
Influence of N-Methylation and Conformation on Almiramide Anti-Leishmanial Activity.Molecules. 2021 Jun 12;26(12):3606. doi: 10.3390/molecules26123606. Molecules. 2021. PMID: 34204673 Free PMC article.
-
Engineering strategy to improve peptide analogs: from structure-based computational design to tumor homing.J Comput Aided Mol Des. 2013 Jan;27(1):31-43. doi: 10.1007/s10822-012-9623-5. Epub 2012 Dec 14. J Comput Aided Mol Des. 2013. PMID: 23239171 Free PMC article.
-
Systematic Exploration of Functional Group Relevance for Anti-Leishmanial Activity of Anisomycin.Biomedicines. 2023 Sep 15;11(9):2541. doi: 10.3390/biomedicines11092541. Biomedicines. 2023. PMID: 37760981 Free PMC article.
-
SwissSidechain: a molecular and structural database of non-natural sidechains.Nucleic Acids Res. 2013 Jan;41(Database issue):D327-32. doi: 10.1093/nar/gks991. Epub 2012 Oct 26. Nucleic Acids Res. 2013. PMID: 23104376 Free PMC article.
References
-
- Voloshchuk N, Montclare JK. Incorporation of unnatural amino acids for synthetic biology. Mol BioSyst. 2010;6:65–80. - PubMed
-
- Wang L, Schultz PG. Expanding the genetic code. Angew Chem Int Ed. 2005;44:34–66. - PubMed
-
- Hendrickson TL, De Crécy-Lagard V, Schimmel P. Incorporation of nonnatural amino acids into proteins. Annu Rev Biochem. 2004;73:147–176. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

