Identification of immunoreactive regions of homology between soluble epidermal growth factor receptor and α5-integrin
- PMID: 21491912
- PMCID: PMC3862712
- DOI: 10.1021/bi200126j
Identification of immunoreactive regions of homology between soluble epidermal growth factor receptor and α5-integrin
Abstract
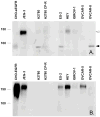
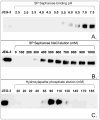
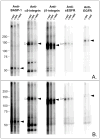
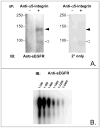
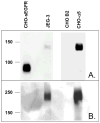
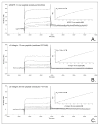
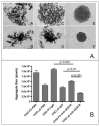
Proteins encoded by the epidermal growth factor receptor (EGFR/HER1/ERBB1) gene are being studied as diagnostic, prognostic, and theragnostic biomarkers for numerous human cancers. The clinical application of these tissue/tumor biomarkers has been limited, in part, by discordant results observed for epidermal growth factor receptor (EGFR) expression using different immunological reagents. Previous studies have used EGFR-directed antibodies that cannot distinguish between full-length and soluble EGFR (sEGFR) expression. We have generated and characterized an anti-sEGFR polyclonal antiserum directed against a 31-mer peptide (residues 604-634) located within the unique 78-amino acid carboxy-terminal sequence of sEGFR. Here, we use this antibody to demonstrate that sEGFR is coexpressed with EGFR in a number of carcinoma-derived cell lines. In addition, we show that a second protein of ~140 kDa (p140) also is detected by this antibody. Rigorous biochemical characterization identifies this second protein to be α5-integrin. We show that a 26-amino acid peptide in the calf domain of α5-integrin (residues 710-735) is 35% identical in sequence with a 31-mer carboxy-terminal sEGFR peptide and exhibits an approximately 5-fold lower affinity for anti-sEGFR than the homologous 31-mer sEGFR peptide does. We conclude that the carboxy terminus of sEGFR and the calf-1 domain of α5-integrin share a region of sequence identity, which results in their mutual immunological reactivity with anti-sEGFR. We also demonstrate that anti-sEGFR promotes three-dimensional tissue cohesion and compaction in vitro, further suggesting a functional link between sEGFR and α5-integrin and a role of the calf-1 domain in cell adhesion. These results have implications for the study of both EGFR and sEGFR as cancer biomarkers and also provide new insight into the mechanisms of interaction between cell surface EGFR isoforms and integrins in complex processes such as cell adhesion and survival signaling.
Figures



Similar articles
-
Shedding of soluble epidermal growth factor receptor (sEGFR) is mediated by a metalloprotease/fibronectin/integrin axis and inhibited by cetuximab.Biochemistry. 2013 Jul 2;52(26):4531-40. doi: 10.1021/bi400437d. Epub 2013 Jun 21. Biochemistry. 2013. PMID: 23731208
-
Integrin α5 Suppresses the Phosphorylation of Epidermal Growth Factor Receptor and Its Cellular Signaling of Cell Proliferation via N-Glycosylation.J Biol Chem. 2015 Dec 4;290(49):29345-60. doi: 10.1074/jbc.M115.682229. Epub 2015 Oct 19. J Biol Chem. 2015. PMID: 26483551 Free PMC article.
-
Anti-EGFR antibody 528 binds to domain III of EGFR at a site shifted from the cetuximab epitope.Sci Rep. 2021 Mar 11;11(1):5790. doi: 10.1038/s41598-021-84171-3. Sci Rep. 2021. PMID: 33707468 Free PMC article.
-
Clinical implementation of soluble EGFR (sEGFR) as a theragnostic serum biomarker of breast, lung and ovarian cancer.IDrugs. 2009 May;12(5):302-8. IDrugs. 2009. PMID: 19431095 Review.
-
Recombinant IgE antibody engineering to target EGFR.Cancer Immunol Immunother. 2012 Sep;61(9):1565-73. doi: 10.1007/s00262-012-1287-4. Epub 2012 Jun 7. Cancer Immunol Immunother. 2012. PMID: 22674055 Free PMC article. Review.
Cited by
-
EGFR soluble isoforms and their transcripts are expressed in meningiomas.PLoS One. 2012;7(5):e37204. doi: 10.1371/journal.pone.0037204. Epub 2012 May 18. PLoS One. 2012. PMID: 22623992 Free PMC article.
-
Multiplex epitope mapping using bacterial surface display reveals both linear and conformational epitopes.Sci Rep. 2012;2:706. doi: 10.1038/srep00706. Epub 2012 Oct 4. Sci Rep. 2012. PMID: 23050090 Free PMC article.
-
EGFR/HER-targeted therapeutics in ovarian cancer.Future Med Chem. 2012 Mar;4(4):447-69. doi: 10.4155/fmc.12.11. Future Med Chem. 2012. PMID: 22416774 Free PMC article. Review.
-
Expression of EGFR isoform D is regulated by HER receptor activators in breast cancer cells.Biochem Biophys Rep. 2022 Aug 19;31:101326. doi: 10.1016/j.bbrep.2022.101326. eCollection 2022 Sep. Biochem Biophys Rep. 2022. PMID: 36039113 Free PMC article.
-
A phase II evaluation of gefitinib in the treatment of persistent or recurrent endometrial cancer: a Gynecologic Oncology Group study.Gynecol Oncol. 2013 Jun;129(3):486-94. doi: 10.1016/j.ygyno.2013.02.019. Epub 2013 Feb 21. Gynecol Oncol. 2013. PMID: 23438670 Free PMC article. Clinical Trial.
References
-
- Yarden Y, Sliwkowski MX. Untangling the ErbB signalling network. Nat Rev Mol Cell Biol. 2001;2:127–137. - PubMed
-
- Boerner JL, Danielsen A, Maihle NJ. Ligand-independent oncogenic signaling by the epidermal growth factor receptor: v-ErbB as a paradigm. Exp Cell Res. 2003;284:111–121. - PubMed
-
- Maihle NJ, Baron AT, Barrette BA, Boardman CH, Christensen TA, Cora EM, Faupel-Badger JM, Greenwood T, Juneja SC, Lafky JM, Lee H, Reiter JL, Podratz KC. EGF/ErbB receptor family in ovarian cancer. Cancer Treat Res. 2002;107:247–258. - PubMed
-
- Reiter JL, Threadgill DW, Eley GD, Strunk KE, Danielsen AJ, Sinclair CS, Pearsall RS, Green PJ, Yee D, Lampland AL, Balasubramaniam S, Crossley TD, Magnuson TR, James CD, Maihle NJ. Comparative genomic sequence analysis and isolation of human and mouse alternative EGFR transcripts encoding truncated receptor isoforms. Genomics. 2001;71:1–20. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

