Coronary artery endothelial transcriptome in vivo: identification of endoplasmic reticulum stress and enhanced reactive oxygen species by gene connectivity network analysis
- PMID: 21493819
- PMCID: PMC3116084
- DOI: 10.1161/CIRCGENETICS.110.958926
Coronary artery endothelial transcriptome in vivo: identification of endoplasmic reticulum stress and enhanced reactive oxygen species by gene connectivity network analysis
Abstract
Background: Endothelial function is central to the localization of atherosclerosis. The in vivo endothelial phenotypic footprints of arterial bed identity and site-specific atherosusceptibility are addressed.
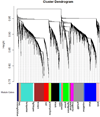
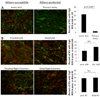
Methods and results: Ninety-eight endothelial cell samples from 13 discrete coronary and noncoronary arterial regions of varying susceptibilities to atherosclerosis were isolated from 76 normal swine. Transcript profiles were analyzed to determine the steady-state in vivo endothelial phenotypes. An unsupervised systems biology approach using weighted gene coexpression networks showed highly correlated endothelial genes. Connectivity network analysis identified 19 gene modules, 12 of which showed significant association with circulatory bed classification. Differential expression of 1300 genes between coronary and noncoronary artery endothelium suggested distinct coronary endothelial phenotypes, with highest significance expressed in gene modules enriched for biological functions related to endoplasmic reticulum (ER) stress and unfolded protein binding, regulation of transcription and translation, and redox homeostasis. Furthermore, within coronary arteries, comparison of endothelial transcript profiles of susceptible proximal regions to protected distal regions suggested the presence of ER stress conditions in susceptible sites. Accumulation of reactive oxygen species throughout coronary endothelium was greater than in noncoronary endothelium consistent with coronary artery ER stress and lower endothelial expression of antioxidant genes in coronary arteries.
Conclusions: Gene connectivity analyses discriminated between coronary and noncoronary endothelial transcript profiles and identified differential transcript levels associated with increased ER and oxidative stress in coronary arteries consistent with enhanced susceptibility to atherosclerosis.
Conflict of interest statement
Figures




References
-
- Schwartz CJ, Mitchell JR. Observations on localization of arterial plaques. Circ Res. 1962;11:63–73. - PubMed
-
- Caro CG, Fitz-Gerald JM, Schroter RC. Arterial wall shear and distribution of early atheroma in man. Nature. 1969;223:1159–1160. - PubMed
-
- McGill HC, Jr, McMahan CA. Determinants of atherosclerosis in the young. Pathobiological determinants of atherosclerosis in youth (PDAY) research group. Am J Cardiol. 1998;82:30T–36T. - PubMed
-
- Gimbrone MA, Jr, Anderson KR, Topper JN, Langille BL, Clowes AW, Bercel S, Davies MG, Stenmark KR, Frid MG, Weiser-Evans MC, Aldashev AA, Nemenoff RA, Majesky MW, Landerholm TE, Lu J, Ito WD, Arras M, Scholz D, Imhof B, Aurrand-Lions M, Schaper W, Nagel TE, Resnick N, Dewey CF, Gimbrone MA, Davies PF. Special communication: the critical role of mechanical forces in blood vessel development, physiology and pathology. J Vasc Surg. 1999;29:1104–1151. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

