Methylation and loss of Secreted Frizzled-Related Protein 3 enhances melanoma cell migration and invasion
- PMID: 21494614
- PMCID: PMC3072980
- DOI: 10.1371/journal.pone.0018674
Methylation and loss of Secreted Frizzled-Related Protein 3 enhances melanoma cell migration and invasion
Abstract
Background: Wnt signaling is important in development and can also contribute to the initiation and progression of cancer. The Secreted Frizzled Related Proteins (SFRPs) constitute a family of Wnt modulators, crucial for controlling Wnt signaling. Here we investigate the expression and role of SFRP3 in melanoma.
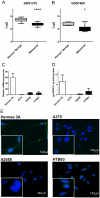
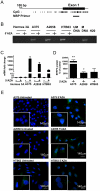
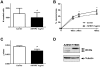
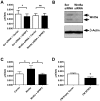
Methodology/principal findings: We show that SFRP3 mRNA is down-regulated in malignant melanoma tumors as compared to normal/benign tissue. Furthermore, we found that SFRP3 expression was lost in the malignant melanoma cell lines, A2058, HTB63 and A375, but not in the non-transformed melanocyte cell line, Hermes 3A. Methylated CpG rich areas were detected in the SFRP3 gene in melanoma cell lines and their SFRP3 expression could be restored using the demethylating agent, 5'aza-deoxycytidine. Addition of recombinant SFRP3 to melanoma cells had no effect on viable cell numbers, but decreased cell migration and invasion. Wnt5a signaling has been shown to increase the migration and invasion of malignant melanoma cells, and high expression of Wnt5a in melanoma tumors has been connected to a poor prognosis. We found that recombinant SFRP3 could inhibit Wnt5a signaling, and that it inhibited melanoma cell migration and invasion in a Wnt5a-dependent manner.
Conclusion/significance: We conclude that SFRP3 functions as a melanoma migration and invasion suppressor by interfering with Wnt5a signaling.
Conflict of interest statement
Figures

Similar articles
-
Methylation-mediated loss of SFRP2 enhances melanoma cell invasion via Wnt signaling.Am J Transl Res. 2016 Mar 15;8(3):1502-9. eCollection 2016. Am J Transl Res. 2016. PMID: 27186276 Free PMC article.
-
A t-butyloxycarbonyl-modified Wnt5a-derived hexapeptide functions as a potent antagonist of Wnt5a-dependent melanoma cell invasion.Proc Natl Acad Sci U S A. 2009 Nov 17;106(46):19473-8. doi: 10.1073/pnas.0909409106. Epub 2009 Nov 9. Proc Natl Acad Sci U S A. 2009. PMID: 19901340 Free PMC article.
-
WNT5A induces release of exosomes containing pro-angiogenic and immunosuppressive factors from malignant melanoma cells.Mol Cancer. 2014 Apr 26;13:88. doi: 10.1186/1476-4598-13-88. Mol Cancer. 2014. PMID: 24766647 Free PMC article.
-
[Crosstalk between Wnt5a and inflammatory signaling in inflammation].Sheng Li Xue Bao. 2015 Aug 25;67(4):437-45. Sheng Li Xue Bao. 2015. PMID: 26300257 Review. Chinese.
-
WNT5A in tumor development and progression: A comprehensive review.Biomed Pharmacother. 2022 Nov;155:113599. doi: 10.1016/j.biopha.2022.113599. Epub 2022 Sep 9. Biomed Pharmacother. 2022. PMID: 36089446 Review.
Cited by
-
Epigenetic loss of putative tumor suppressor SFRP3 correlates with poor prognosis of lung adenocarcinoma patients.Epigenetics. 2018;13(3):214-227. doi: 10.1080/15592294.2016.1229730. Epub 2018 Apr 18. Epigenetics. 2018. PMID: 27623992 Free PMC article.
-
Low expression of RBMS3 and SFRP1 are associated with poor prognosis in patients with gastric cancer.Am J Cancer Res. 2016 Nov 1;6(11):2679-2689. eCollection 2016. Am J Cancer Res. 2016. PMID: 27904780 Free PMC article.
-
Wnt signaling in cancer.Cold Spring Harb Perspect Biol. 2012 May 1;4(5):a008052. doi: 10.1101/cshperspect.a008052. Cold Spring Harb Perspect Biol. 2012. PMID: 22438566 Free PMC article. Review.
-
A transcriptome-based protein network that identifies new therapeutic targets in colorectal cancer.BMC Genomics. 2017 Sep 30;18(1):758. doi: 10.1186/s12864-017-4139-y. BMC Genomics. 2017. PMID: 28962550 Free PMC article.
-
Integrated analysis of genome-wide DNA methylation and gene expression profiles identifies potential novel biomarkers of rectal cancer.Oncotarget. 2016 Sep 20;7(38):62547-62558. doi: 10.18632/oncotarget.11534. Oncotarget. 2016. PMID: 27566576 Free PMC article.
References
-
- Rao TP, Kuhl M. An updated overview on Wnt signaling pathways: a prelude for more. Circ Res. 2010;106:1798–1806. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

