Modulating repeat protein stability: the effect of individual helix stability on the collective behavior of the ensemble
- PMID: 21495096
- PMCID: PMC3104233
- DOI: 10.1002/pro.638
Modulating repeat protein stability: the effect of individual helix stability on the collective behavior of the ensemble
Abstract
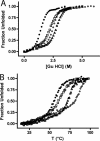
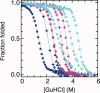
Repeat proteins are tandem arrays of a small structural motif, in which tertiary structure is stabilized by interactions within a repeat and between neighboring repeats. Several studies have shown that this modular structure is manifest in modular thermodynamic properties. Specifically, the global stability of a repeat protein can be described by simple linear models, considering only two parameters: the stability of the individual repeated units (H) and the coupling interaction between the units (J). If the repeat units are identical, single values of H and J, together with the number of repeated units, is sufficient to completely describe the thermodynamic behavior of any protein within a series. In this work, we demonstrate how the global stability of a repeat protein can be changed, in a predictable fashion, by modifying only the H parameter. Taking a previously characterized series of consensus tetratricopeptide repeats (TPR) (CTPRa) proteins, we introduced mutations into the basic repeating unit, such that the stability of the individual repeat unit was increased, but its interaction with neighboring units was unchanged. In other words, we increased H but kept J constant. We demonstrated that the denaturation curves for a series of such repeat proteins can be fit and additional curves can be predicted by the one-dimensional Ising model in which only H has changed from the original fit for the CTPRa series. Our results show that we can significantly increase the stability of a repeat protein by rationally increasing the stability of the units (H), whereas the interaction between repeats (J) remains unchanged.
Copyright © 2011 The Protein Society.
Figures

Similar articles
-
Calorimetric study of a series of designed repeat proteins: modular structure and modular folding.Protein Sci. 2011 Feb;20(2):336-40. doi: 10.1002/pro.564. Protein Sci. 2011. PMID: 21280125 Free PMC article.
-
Exploring the folding energy landscape of a series of designed consensus tetratricopeptide repeat proteins.Proc Natl Acad Sci U S A. 2009 Oct 13;106(41):17383-8. doi: 10.1073/pnas.0907455106. Epub 2009 Oct 1. Proc Natl Acad Sci U S A. 2009. PMID: 19805120 Free PMC article.
-
Local and long-range stability in tandemly arrayed tetratricopeptide repeats.Proc Natl Acad Sci U S A. 2005 Apr 19;102(16):5721-6. doi: 10.1073/pnas.0404530102. Epub 2005 Apr 11. Proc Natl Acad Sci U S A. 2005. PMID: 15824314 Free PMC article.
-
Repeat proteins challenge the concept of structural domains.Biochem Soc Trans. 2015 Oct;43(5):844-9. doi: 10.1042/BST20150083. Biochem Soc Trans. 2015. PMID: 26517892 Review.
-
The tetratricopeptide repeat: a structural motif mediating protein-protein interactions.Bioessays. 1999 Nov;21(11):932-9. doi: 10.1002/(SICI)1521-1878(199911)21:11<932::AID-BIES5>3.0.CO;2-N. Bioessays. 1999. PMID: 10517866 Review.
Cited by
-
Exploring new strategies for grafting binding peptides onto protein loops using a consensus-designed tetratricopeptide repeat scaffold.Protein Sci. 2019 Apr;28(4):738-745. doi: 10.1002/pro.3586. Protein Sci. 2019. PMID: 30746804 Free PMC article.
-
A designed repeat protein as an affinity capture reagent.Biochem Soc Trans. 2015 Oct;43(5):874-80. doi: 10.1042/BST20150091. Biochem Soc Trans. 2015. PMID: 26517897 Free PMC article. Review.
-
Removal of a consensus proline is not sufficient to allow tetratricopeptide repeat oligomerization.Protein Sci. 2017 Oct;26(10):1974-1983. doi: 10.1002/pro.3234. Epub 2017 Jul 25. Protein Sci. 2017. PMID: 28707340 Free PMC article.
-
Nanostructured functional films from engineered repeat proteins.J R Soc Interface. 2013 Apr 17;10(83):20130051. doi: 10.1098/rsif.2013.0051. Print 2013 Jun 6. J R Soc Interface. 2013. PMID: 23594813 Free PMC article.
-
Engineering bio-brick protein scaffolds for organizing enzyme assemblies.Protein Sci. 2024 May;33(5):e4984. doi: 10.1002/pro.4984. Protein Sci. 2024. PMID: 38607190 Free PMC article.
References
-
- Andrade MA, Perez-Iratxeta C, Ponting CP. Protein repeats: structures, functions, and evolution. J Struct Biol. 2001;134:117–131. - PubMed
-
- Groves MR, Barford D. Topological characteristics of helical repeat proteins. Curr Opin Struct Biol. 1999;9:383–389. - PubMed
-
- Main E, Lowe A, Mochrie S, Jackson S, Regan L. A recurring theme in protein engineering: the design, stability and folding of repeat proteins. Curr Opin Struct Biol. 2005;15:464–471. - PubMed
-
- D'Andrea L, Regan L. TPR proteins: the versatile helix. Trends Biochem Sci. 2003;28:655–662. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

