Structure of Francisella tularensis peptidyl-tRNA hydrolase
- PMID: 21505237
- PMCID: PMC3080146
- DOI: 10.1107/S174430911100515X
Structure of Francisella tularensis peptidyl-tRNA hydrolase
Abstract
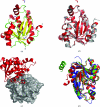
The rational design of novel antibiotics for bacteria involves the identification of inhibitors for enzymes involved in essential biochemical pathways in cells. In this study, the cloning, expression, purification, crystallization and structure of the enzyme peptidyl-tRNA hydrolase from Francisella tularensis, the causative agent of tularemia, was performed. The structure of F. tularensis peptidyl-tRNA hydrolase is comparable to those of other bacterial peptidyl-tRNA hydrolases, with most residues in the active site conserved amongst the family. The resultant reagents, structural data and analyses provide essential information for the structure-based design of novel inhibitors for this class of proteins.
Figures
Similar articles
-
Crystal structure at 1.2 A resolution and active site mapping of Escherichia coli peptidyl-tRNA hydrolase.EMBO J. 1997 Aug 1;16(15):4760-9. doi: 10.1093/emboj/16.15.4760. EMBO J. 1997. PMID: 9303320 Free PMC article.
-
The mode of inhibitor binding to peptidyl-tRNA hydrolase: binding studies and structure determination of unbound and bound peptidyl-tRNA hydrolase from Acinetobacter baumannii.PLoS One. 2013 Jul 3;8(7):e67547. doi: 10.1371/journal.pone.0067547. Print 2013. PLoS One. 2013. PMID: 23844024 Free PMC article.
-
Cloning, expression, purification, crystallization and preliminary X-ray analysis of peptidyl-tRNA hydrolase from Mycobacterium tuberculosis.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2006 Sep 1;62(Pt 9):913-5. doi: 10.1107/S1744309106031125. Epub 2006 Aug 26. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2006. PMID: 16946478 Free PMC article.
-
Structural and functional insights into peptidyl-tRNA hydrolase.Biochim Biophys Acta. 2014 Jul;1844(7):1279-88. doi: 10.1016/j.bbapap.2014.04.012. Epub 2014 Apr 21. Biochim Biophys Acta. 2014. PMID: 24768774 Review.
-
Unveiling the Druggable Landscape of Bacterial Peptidyl tRNA Hydrolase: Insights into Structure, Function, and Therapeutic Potential.Biomolecules. 2024 Jun 7;14(6):668. doi: 10.3390/biom14060668. Biomolecules. 2024. PMID: 38927071 Free PMC article. Review.
Cited by
-
Crystallization and preliminary X-ray analysis of peptidyl-tRNA hydrolase from Thermus thermophilus HB8.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2013 Mar 1;69(Pt 3):332-5. doi: 10.1107/S1744309113003424. Epub 2013 Feb 27. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2013. PMID: 23519816 Free PMC article.
-
Small Molecule Docking Supports Broad and Narrow Spectrum Potential for the Inhibition of the Novel Antibiotic Target Bacterial Pth1.Antibiotics (Basel). 2016 May 10;5(2):16. doi: 10.3390/antibiotics5020016. Antibiotics (Basel). 2016. PMID: 27171117 Free PMC article.
-
Crystal structure of peptidyl-tRNA hydrolase from a Gram-positive bacterium, Streptococcus pyogenes at 2.19 Å resolution shows the closed structure of the substrate-binding cleft.FEBS Open Bio. 2014 Oct 22;4:915-22. doi: 10.1016/j.fob.2014.10.010. eCollection 2014. FEBS Open Bio. 2014. PMID: 25389518 Free PMC article.
-
Recombinant production, crystallization and X-ray crystallographic structure determination of the peptidyl-tRNA hydrolase of Pseudomonas aeruginosa.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2012 Dec 1;68(Pt 12):1472-6. doi: 10.1107/S1744309112045770. Epub 2012 Nov 28. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2012. PMID: 23192026 Free PMC article.
-
Structural basis for the substrate recognition and catalysis of peptidyl-tRNA hydrolase.Nucleic Acids Res. 2012 Nov 1;40(20):10521-31. doi: 10.1093/nar/gks790. Epub 2012 Aug 25. Nucleic Acids Res. 2012. PMID: 22923517 Free PMC article.
References
-
- Brun, G., Paulin, D., Yot, P. & Chapeville, F. (1971). Biochimie, 53, 225–231. - PubMed
-
- Collaborative Computational Project, Number 4 (1994). Acta Cryst. D50, 760–763. - PubMed
-
- DeLano, W. L. (2002). PyMOL. http://www.pymol.org.
-
- De Pereda, J. M., Waas, W. F., Jan, Y., Ruoslahti, E., Schimmel, P. & Pascual, J. (2004). J. Biol. Chem. 279, 8111–8115. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources

