Sulfatases and a radical S-adenosyl-L-methionine (AdoMet) enzyme are key for mucosal foraging and fitness of the prominent human gut symbiont, Bacteroides thetaiotaomicron
- PMID: 21507958
- PMCID: PMC3138274
- DOI: 10.1074/jbc.M111.228841
Sulfatases and a radical S-adenosyl-L-methionine (AdoMet) enzyme are key for mucosal foraging and fitness of the prominent human gut symbiont, Bacteroides thetaiotaomicron
Abstract
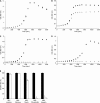
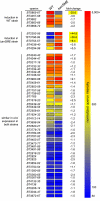
The large-scale application of genomic and metagenomic sequencing technologies has yielded a number of insights about the metabolic potential of symbiotic human gut microbes. Nevertheless, the molecular basis of the interactions between commensal bacteria and their host remained to be investigated. Bacteria colonizing the mucosal layer that overlies the gut epithelium are exposed to highly sulfated glycans (i.e. mucin and glycosaminoglycans). These polymers can serve as potential nutrient sources, but their high sulfate content usually prevents their degradation. Commensal bacteria such as Bacteroides thetaiotaomicron possess more predicted sulfatase genes than in the human genome, the physiological functions of which are largely unknown. To be active, sulfatases must undergo a critical post-translational modification catalyzed in anaerobic bacteria by the radical AdoMet enzyme anaerobic sulfatase-maturating enzyme (anSME). In the present study, we have tested the role of this pathway in Bacteroides thetaiotaomicron which, in addition to 28 predicted sulfatases, possesses a single predicted anSME. In vitro studies revealed that deletion of the gene encoding its anSME (BT0238) results in loss of sulfatase activity and impaired ability to use sulfated polysaccharides as carbon sources. Co-colonization of formerly germ-free mice with both isogenic strains (i.e. wild-type or ΔanSME), or invasion experiments involving introduction of one followed by the other strain established that anSME activity and the sulfatases activated via this pathway, are important fitness factors for B. thetaiotaomicron, especially when mice are fed a simple sugar diet that requires this saccharolytic bacterium to adaptively forage on host glycans as nutrients. Whole genome transcriptional profiling of wild-type and the anSME mutant in vivo revealed that loss of this enzyme alters expression of genes involved in mucin utilization and that this disrupted ability to access mucosal glycans likely underlies the observed pronounced colonization defect. Comparative genomic analysis reveals that 100% of 46 fully sequenced human gut Bacteroidetes contain homologs of BT0238 and genes encoding sulfatases, suggesting that this is an important and evolutionarily conserved feature for bacterial adaptation to life in this habitat.
Figures





References
-
- Qin J., Li R., Raes J., Arumugam M., Burgdorf K. S., Manichanh C., Nielsen T., Pons N., Levenez F., Yamada T., Mende D. R., Li J., Xu J., Li S., Li D., Cao J., Wang B., Liang H., Zheng H., Xie Y., Tap J., Lepage P., Bertalan M., Batto J. M., Hansen T., Le Paslier D., Linneberg A., Nielsen H. B., Pelletier E., Renault P., Sicheritz-Ponten T., Turner K., Zhu H., Yu C., Li S., Jian M., Zhou Y., Li Y., Zhang X., Li S., Qin N., Yang H., Wang J., Brunak S., Doré J., Guarner F., Kristiansen K., Pedersen O., Parkhill J., Weissenbach J., Bork P., Ehrlich S. D., Wang J. (2010) Nature 464, 59–65 - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

