A survey of metabolic databases emphasizing the MetaCyc family
- PMID: 21523460
- PMCID: PMC3352032
- DOI: 10.1007/s00204-011-0705-2
A survey of metabolic databases emphasizing the MetaCyc family
Abstract
Thanks to the confluence of genome sequencing and bioinformatics, the number of metabolic databases has expanded from a handful in the mid-1990s to several thousand today. These databases lie within distinct families that have common ancestry and common attributes. The main families are the MetaCyc, KEGG, Reactome, Model SEED, and BiGG families. We survey these database families, as well as important individual metabolic databases, including multiple human metabolic databases. The MetaCyc family is described in particular detail. It contains well over 1,000 databases, including highly curated databases for Escherichia coli, Saccharomyces cerevisiae, Mus musculus, and Arabidopsis thaliana. These databases are available through a number of web sites that offer a range of software tools for querying and visualizing metabolic networks. These web sites also provide multiple tools for analysis of gene expression and metabolomics data, including visualization of those datasets on metabolic network diagrams and over-representation analysis of gene sets and metabolite sets.
Figures
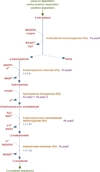











References
-
- Bernal V, Carinhas N, Yokomizo AY, Carrondo MJ, Alves PM. Cell density effect in the baculovirus-insect cells system: a quantitative analysis of energetic metabolism. Biotechnol Bioeng. 2009;104(1):162–180. - PubMed
-
- BioCyc webinars. SRI International; http://biocyc.org/webinar.shtml,
-
- Caspi R, Altman T, Dale JM, Dreher K, Fulcher CA, Gilham F, Kaipa P, Karthikeyan AS, Kothari A, Krummenacker M, Latendresse M, Mueller LA, Paley S, Popescu L, Pujar A, Shearer AG, Zhang P, Karp PD. The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of pathway/genome databases. Nucleic Acids Res. 2010;38(Database issue):D473–D479. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

