Mitochondrial protein turnover: role of the precursor intermediate peptidase Oct1 in protein stabilization
- PMID: 21525245
- PMCID: PMC3128517
- DOI: 10.1091/mbc.E11-02-0169
Mitochondrial protein turnover: role of the precursor intermediate peptidase Oct1 in protein stabilization
Abstract
Most mitochondrial proteins are encoded in the nucleus as precursor proteins and carry N-terminal presequences for import into the organelle. The vast majority of presequences are proteolytically removed by the mitochondrial processing peptidase (MPP) localized in the matrix. A subset of precursors with a characteristic amino acid motif is additionally processed by the mitochondrial intermediate peptidase (MIP) octapeptidyl aminopeptidase 1 (Oct1), which removes an octapeptide from the N-terminus of the precursor intermediate. However, the function of this second cleavage step is elusive. In this paper, we report the identification of a novel Oct1 substrate protein with an unusual cleavage motif. Inspection of the Oct1 substrates revealed that the N-termini of the intermediates typically carry a destabilizing amino acid residue according to the N-end rule of protein degradation, whereas mature proteins carry stabilizing N-terminal residues. We compared the stability of intermediate and mature forms of Oct1 substrate proteins in organello and in vivo and found that Oct1 cleavage increases the half-life of its substrate proteins, most likely by removing destabilizing amino acids at the intermediate's N-terminus. Thus Oct1 converts unstable precursor intermediates generated by MPP into stable mature proteins.
Figures

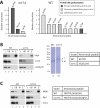



Similar articles
-
More than just a ticket canceller: the mitochondrial processing peptidase tailors complex precursor proteins at internal cleavage sites.Mol Biol Cell. 2020 Nov 15;31(24):2657-2668. doi: 10.1091/mbc.E20-08-0524. Epub 2020 Sep 30. Mol Biol Cell. 2020. PMID: 32997570 Free PMC article.
-
Prediction and identification of new natural substrates of the yeast mitochondrial intermediate peptidase.J Biol Chem. 1995 Nov 10;270(45):27366-73. doi: 10.1074/jbc.270.45.27366. J Biol Chem. 1995. PMID: 7593000
-
Amino-terminal octapeptides function as recognition signals for the mitochondrial intermediate peptidase.J Biol Chem. 1992 Apr 15;267(11):7904-10. J Biol Chem. 1992. PMID: 1560019
-
Mitochondrial processing peptidases.Biochim Biophys Acta. 2002 Sep 2;1592(1):63-77. doi: 10.1016/s0167-4889(02)00265-3. Biochim Biophys Acta. 2002. PMID: 12191769 Review.
-
Processing of mitochondrial precursor proteins.Biomed Biochim Acta. 1991;50(4-6):403-12. Biomed Biochim Acta. 1991. PMID: 1839352 Review.
Cited by
-
More than just a ticket canceller: the mitochondrial processing peptidase tailors complex precursor proteins at internal cleavage sites.Mol Biol Cell. 2020 Nov 15;31(24):2657-2668. doi: 10.1091/mbc.E20-08-0524. Epub 2020 Sep 30. Mol Biol Cell. 2020. PMID: 32997570 Free PMC article.
-
INTERMEDIATE CLEAVAGE PEPTIDASE55 Modifies Enzyme Amino Termini and Alters Protein Stability in Arabidopsis Mitochondria.Plant Physiol. 2015 Jun;168(2):415-27. doi: 10.1104/pp.15.00300. Epub 2015 Apr 10. Plant Physiol. 2015. PMID: 25862457 Free PMC article.
-
Role of the Mitochondrial Protein Import Machinery and Protein Processing in Heart Disease.Front Cardiovasc Med. 2021 Sep 28;8:749756. doi: 10.3389/fcvm.2021.749756. eCollection 2021. Front Cardiovasc Med. 2021. PMID: 34651031 Free PMC article. Review.
-
Human mitochondrial peroxiredoxin Prdx3 is dually localized in the intermembrane space and matrix subcompartments.Redox Biol. 2024 Dec;78:103436. doi: 10.1016/j.redox.2024.103436. Epub 2024 Nov 21. Redox Biol. 2024. PMID: 39591905 Free PMC article.
-
Identification of cleavage sites and substrate proteins for two mitochondrial intermediate peptidases in Arabidopsis thaliana.J Exp Bot. 2015 May;66(9):2691-708. doi: 10.1093/jxb/erv064. Epub 2015 Mar 1. J Exp Bot. 2015. PMID: 25732537 Free PMC article.
References
-
- Bachmair A, Finley D, Varshavsky A. In vivo half-life of a protein is a function of its amino-terminal residue. Science. 1986;234:179–186. - PubMed
-
- Beckmann JD, Ljungdahl PO, Lopez JL, Trumpower BL. Isolation and characterization of the nuclear gene encoding the Rieske iron-sulfur protein (RIP1) from Saccharomyces cerevisiae. J Biol Chem. 1987;262:8901–8909. - PubMed
-
- Branda SS, Isaya G. Prediction and identification of new natural substrates of the yeast mitochondrial intermediate peptidase. J Biol Chem. 1995;270:27366–27373. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

