Origin of bistability underlying mammalian cell cycle entry
- PMID: 21525871
- PMCID: PMC3101952
- DOI: 10.1038/msb.2011.19
Origin of bistability underlying mammalian cell cycle entry
Abstract
Precise control of cell proliferation is fundamental to tissue homeostasis and differentiation. Mammalian cells commit to proliferation at the restriction point (R-point). It has long been recognized that the R-point is tightly regulated by the Rb-E2F signaling pathway. Our recent work has further demonstrated that this regulation is mediated by a bistable switch mechanism. Nevertheless, the essential regulatory features in the Rb-E2F pathway that create this switching property have not been defined. Here we analyzed a library of gene circuits comprising all possible link combinations in a simplified Rb-E2F network. We identified a minimal circuit that is able to generate robust, resettable bistability. This minimal circuit contains a feed-forward loop coupled with a mutual-inhibition feedback loop, which forms an AND-gate control of the E2F activation. Underscoring its importance, experimental disruption of this circuit abolishes maintenance of the activated E2F state, supporting its importance for the bistability of the Rb-E2F system. Our findings suggested basic design principles for the robust control of the bistable cell cycle entry at the R-point.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
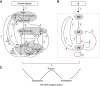
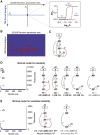


References
-
- Alon U, Surette MG, Barkai N, Leibler S (1999) Robustness in bacterial chemotaxis. Nature 397: 168–171 - PubMed
-
- Barkai N, Leibler S (1997) Robustness in simple biochemical networks. Nature 387: 913–917 - PubMed
-
- Bartek J, Bartkova J, Lukas J (1996) The retinoblastoma protein pathway and the restriction point. Curr Opin Cell Biol 8: 805–814 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

