Identification of potential conserved RNA secondary structure throughout influenza A coding regions
- PMID: 21536710
- PMCID: PMC3096049
- DOI: 10.1261/rna.2619511
Identification of potential conserved RNA secondary structure throughout influenza A coding regions
Abstract
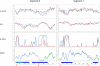
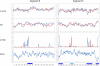
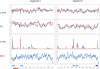
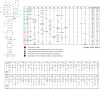
Influenza A is a negative sense RNA virus of significant public health concern. While much is understood about the life cycle of the virus, knowledge of RNA secondary structure in influenza A virus is sparse. Predictions of RNA secondary structure can focus experimental efforts. The present study analyzes coding regions of the eight viral genome segments in both the (+) and (-) sense RNA for conserved secondary structure. The predictions are based on identifying regions of unusual thermodynamic stabilities and are correlated with studies of suppression of synonymous codon usage (SSCU). The results indicate that secondary structure is favored in the (+) sense influenza RNA. Twenty regions with putative conserved RNA structure have been identified, including two previously described structured regions. Of these predictions, eight have high thermodynamic stability and SSCU, with five of these corresponding to current annotations (e.g., splice sites), while the remaining 12 are predicted by the thermodynamics alone. Secondary structures with high conservation of base-pairing are proposed within the five regions having known function. A combination of thermodynamics, amino acid and nucleotide sequence comparisons along with SSCU was essential for revealing potential secondary structures.
Figures



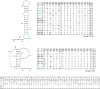




References
-
- Basler CF, Reid AH, Dybing JK, Janczewski TA, Fanning TG, Zheng H, Salvatore M, Perdue ML, Swayne DE, García-Sastre A 2001. Sequence of the 1918 pandemic influenza virus nonstructural gene (NS) segment and characterization of recombinant viruses bearing the 1918 NS genes. Proc Natl Acad Sci 98: 2746–2751 - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
